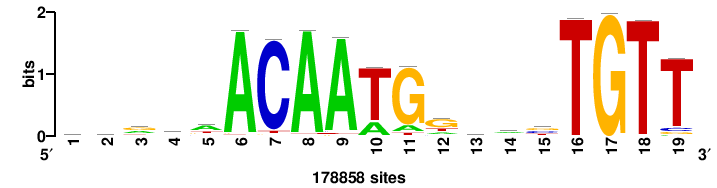
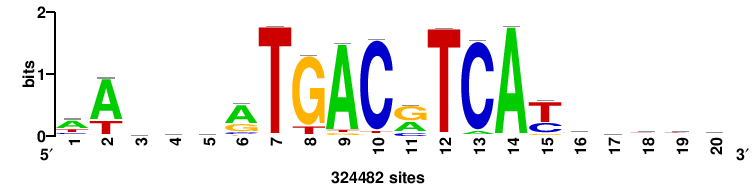
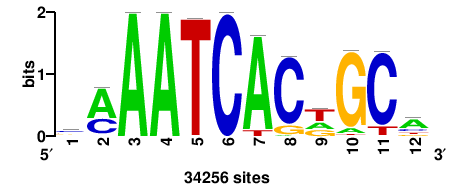
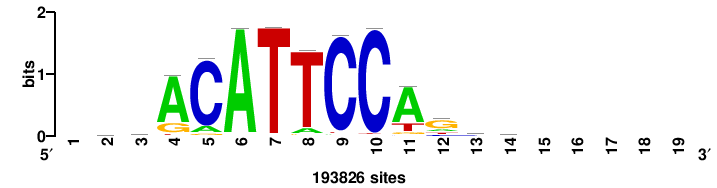
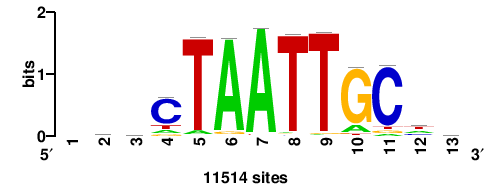
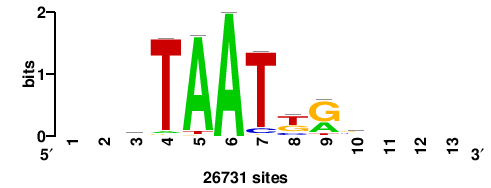
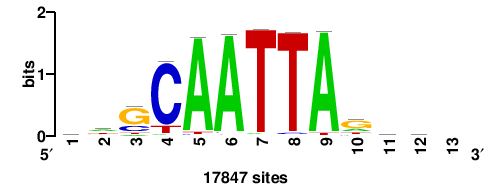
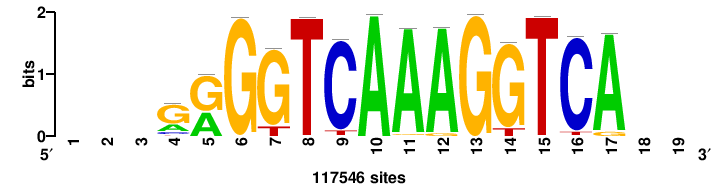
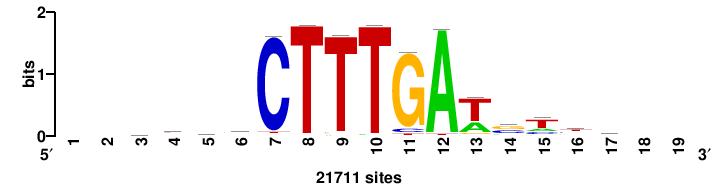
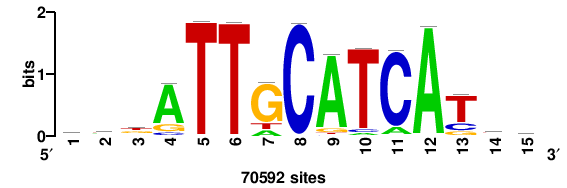
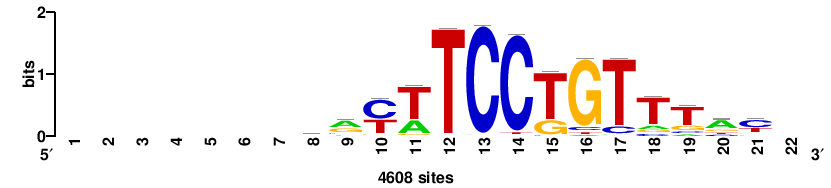
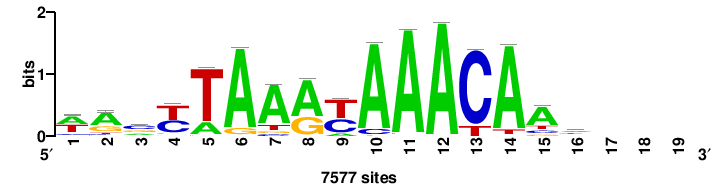
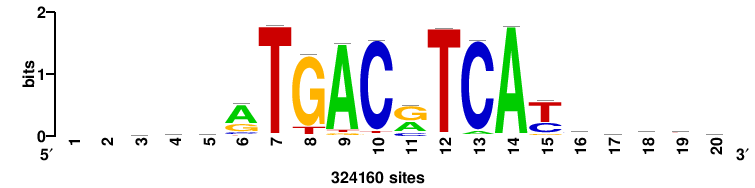
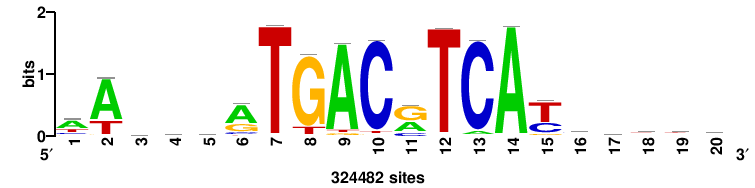
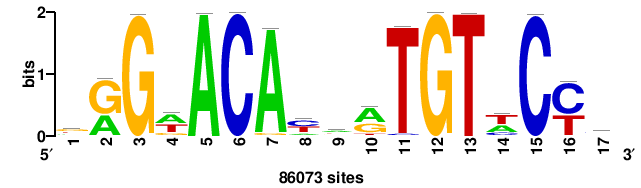
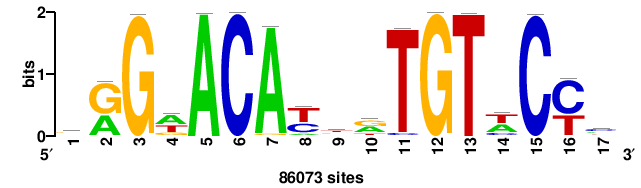
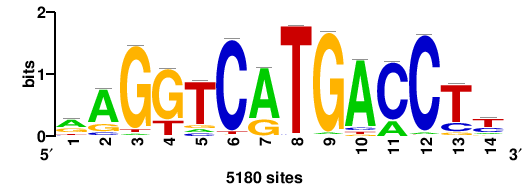
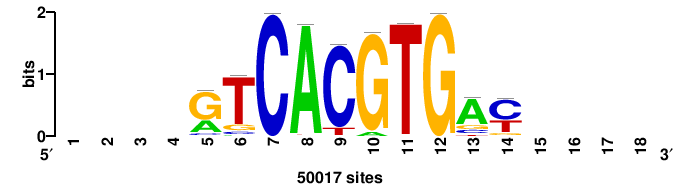
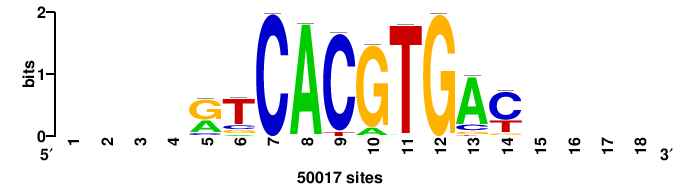
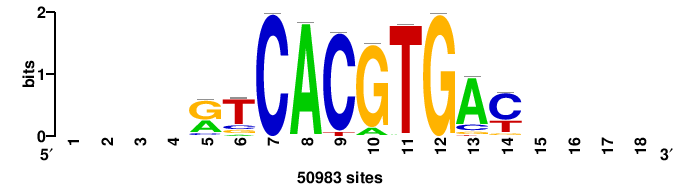
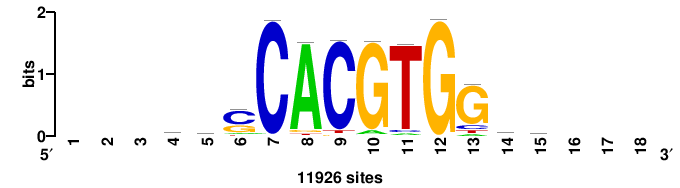
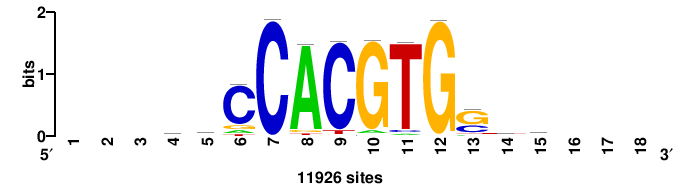
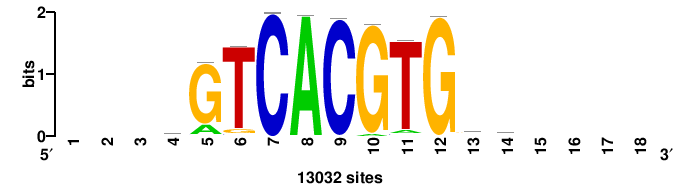
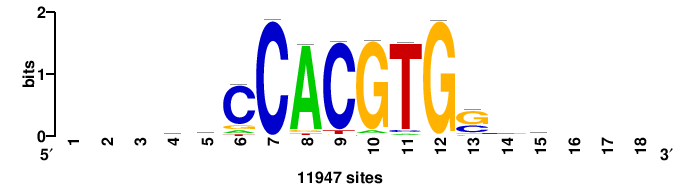
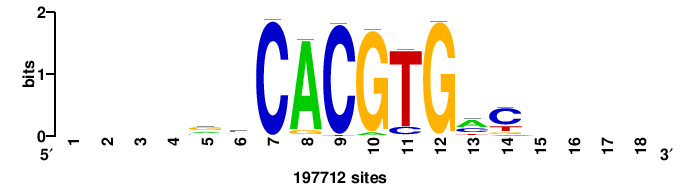
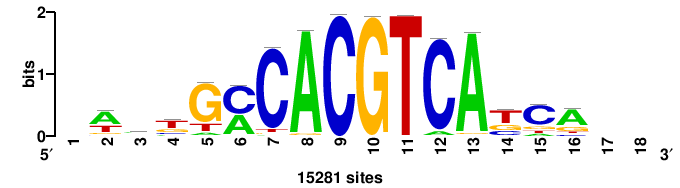
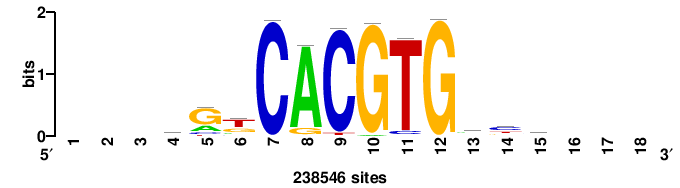
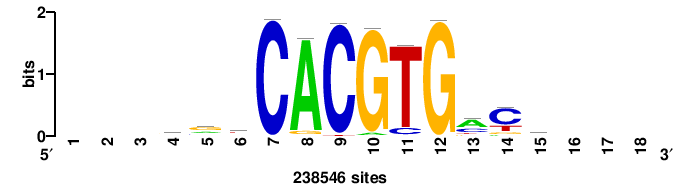
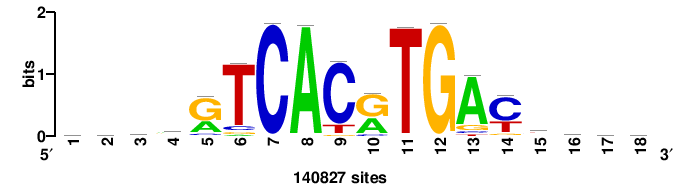
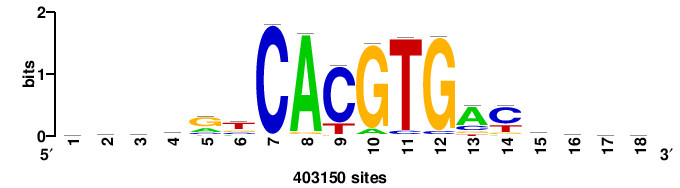
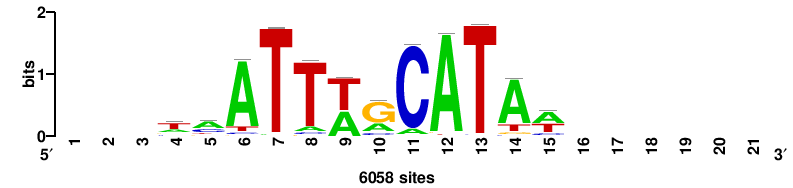
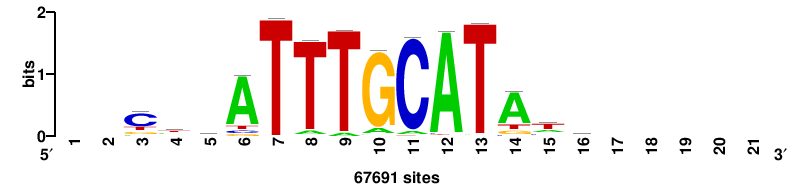
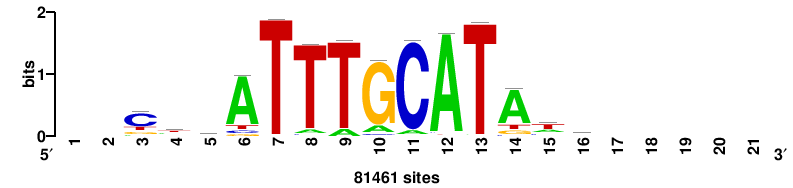
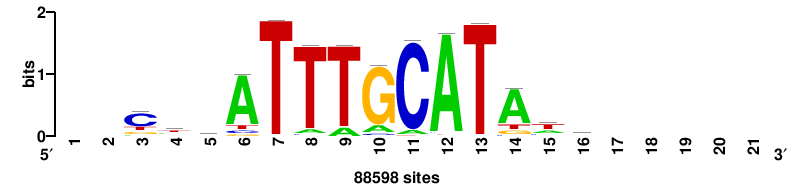
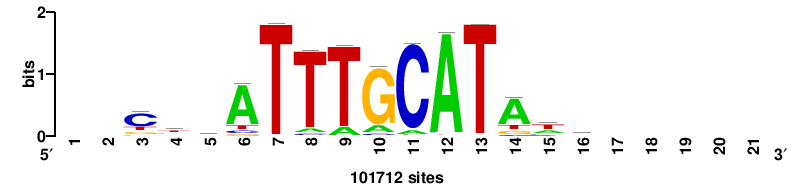
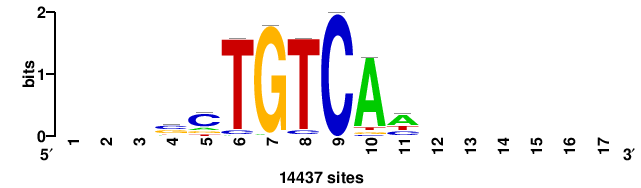
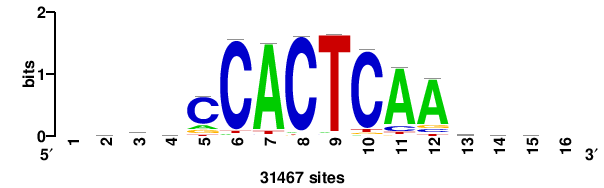
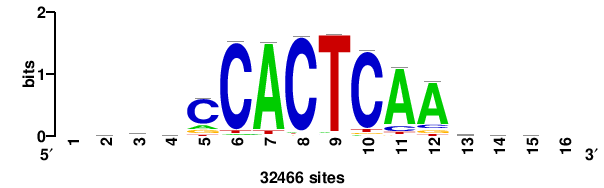
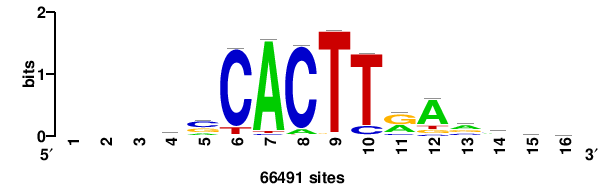
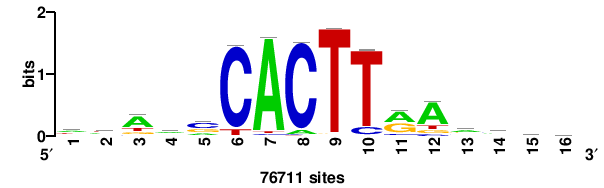
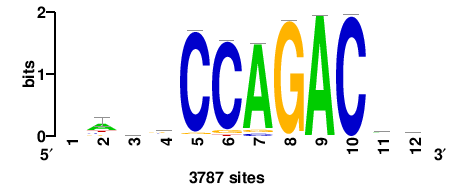
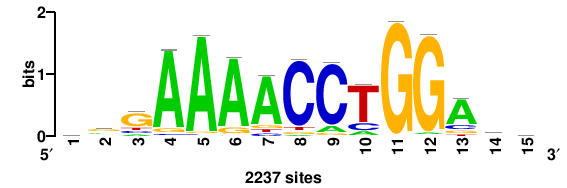
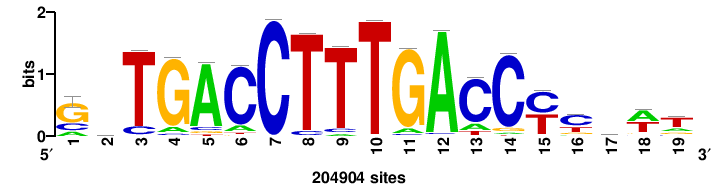
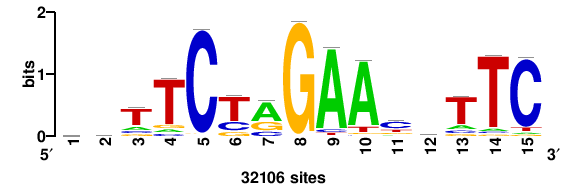
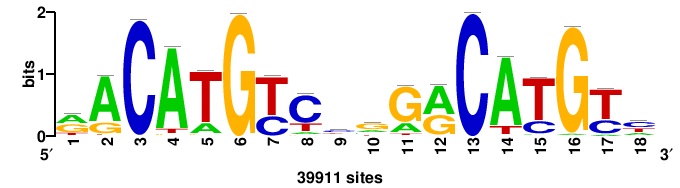
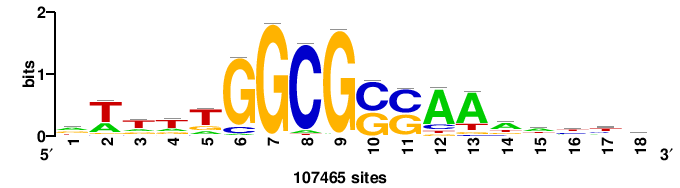
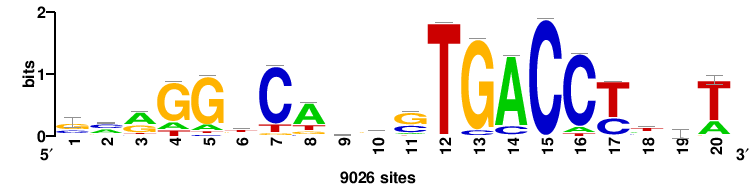
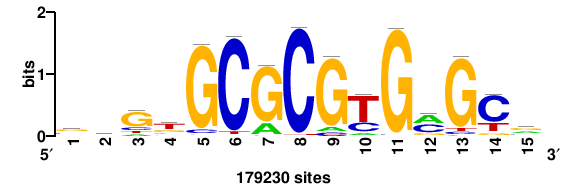
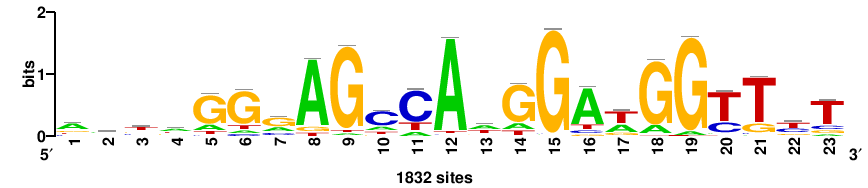
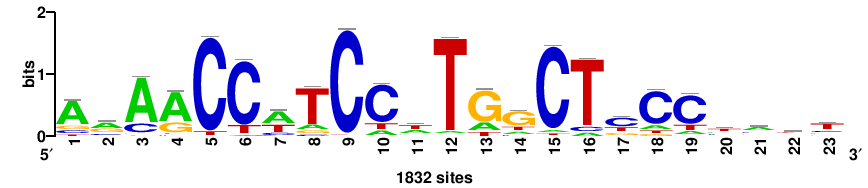
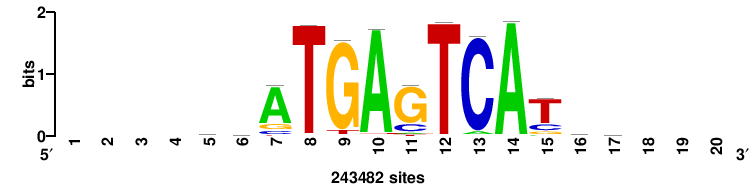
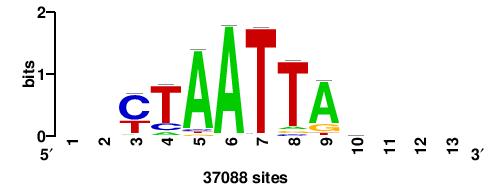
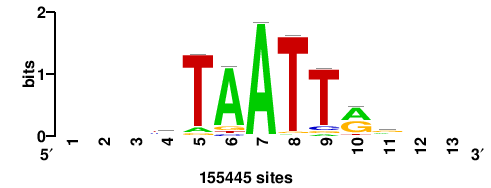
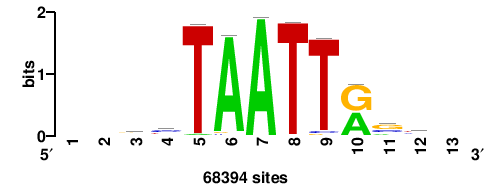
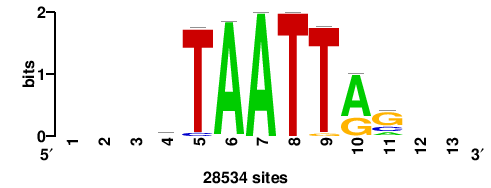
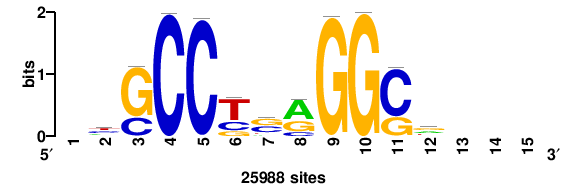
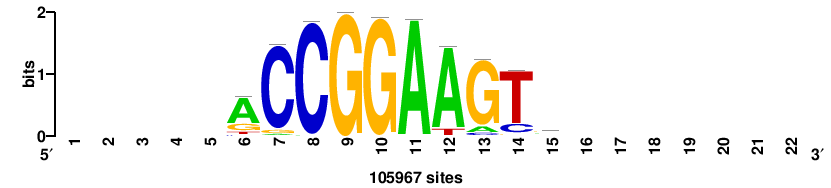
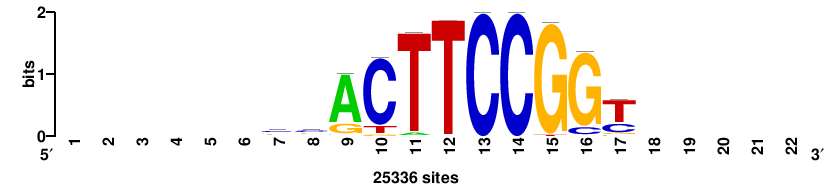
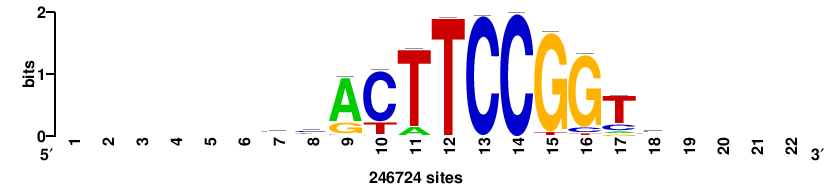
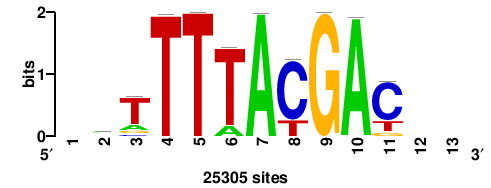
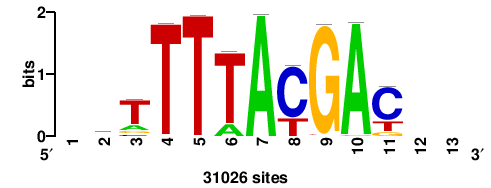
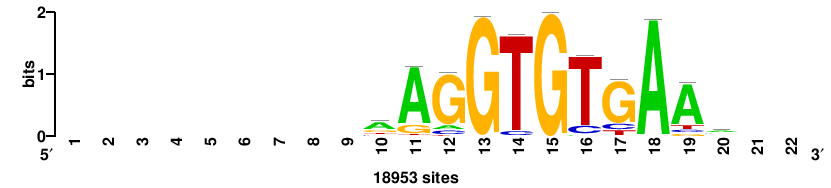
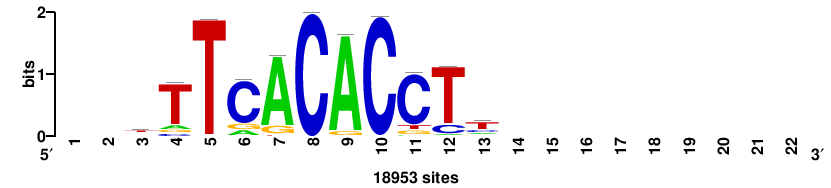
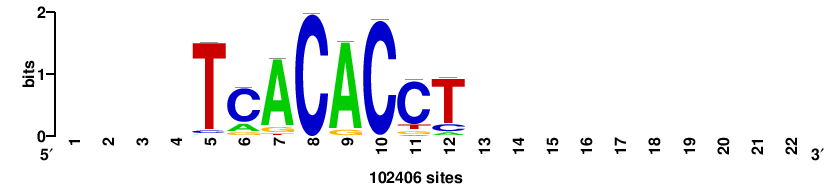
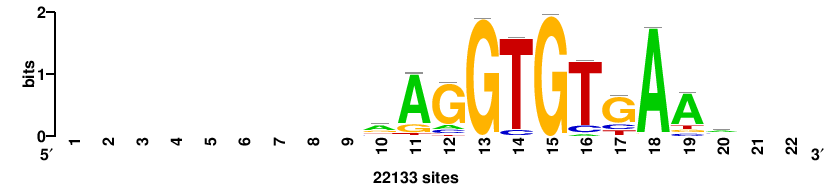
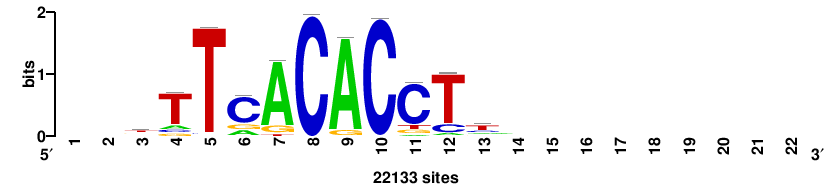
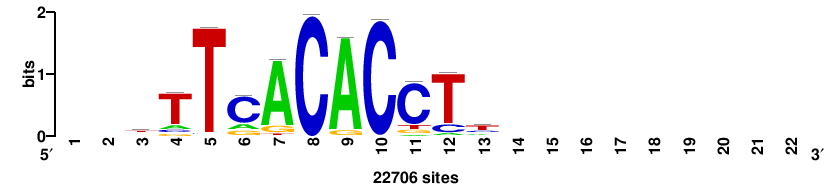
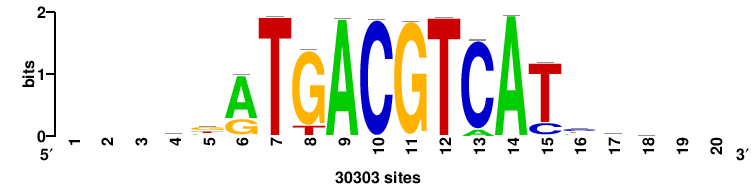
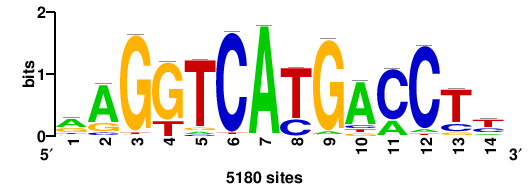
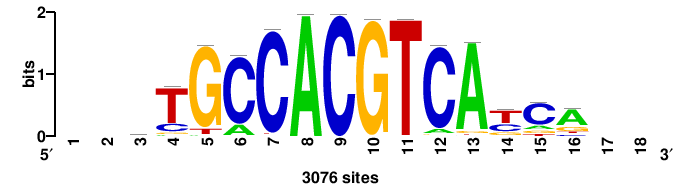
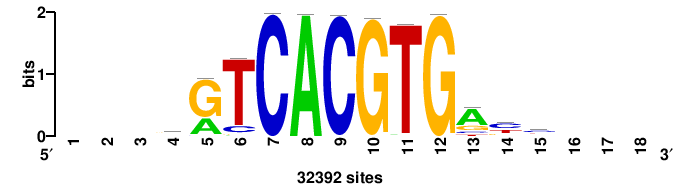
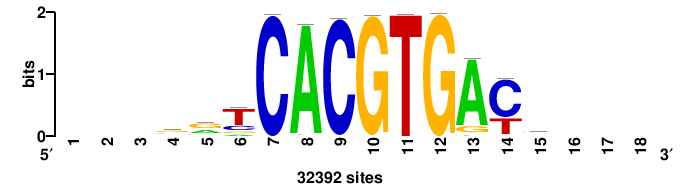
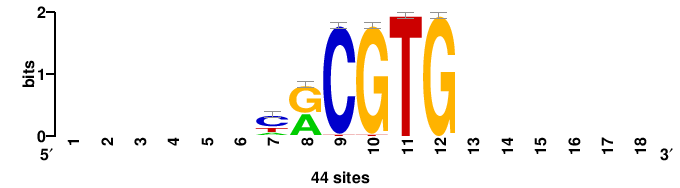
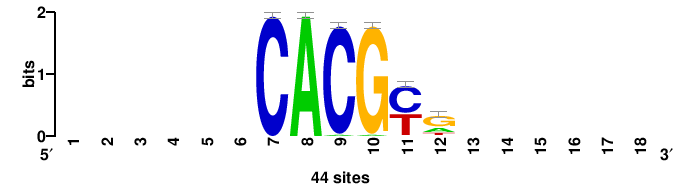
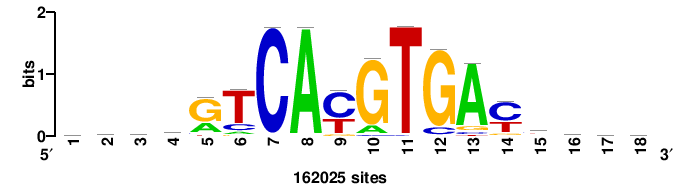
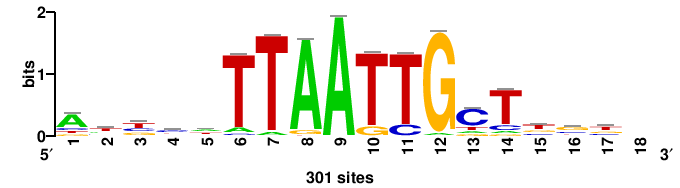
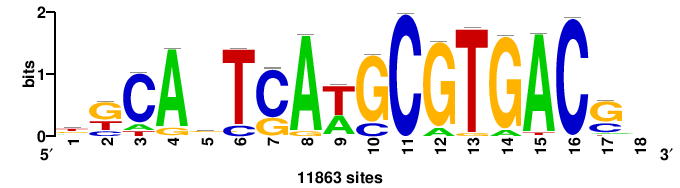
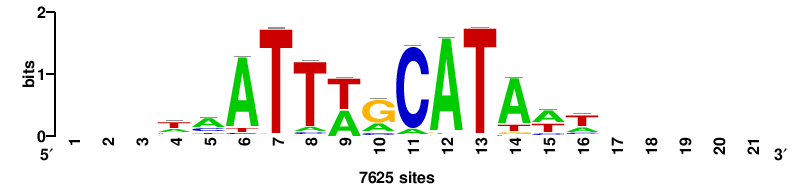
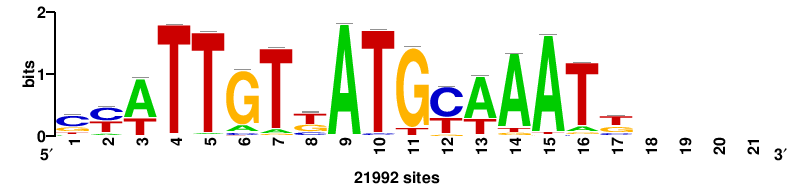
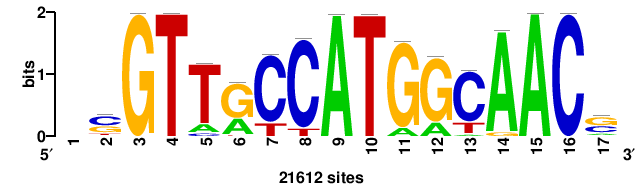
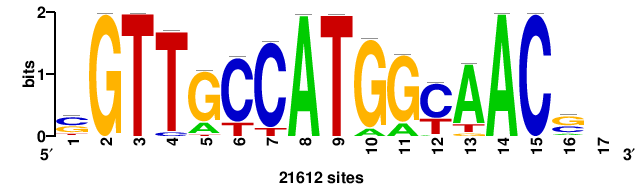
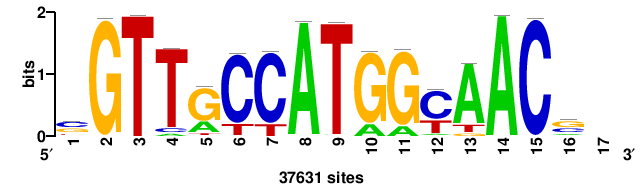
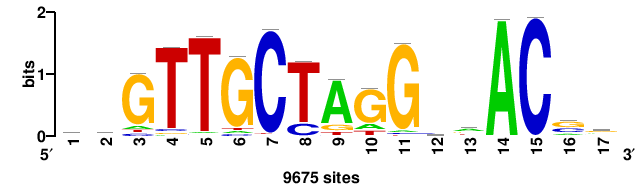
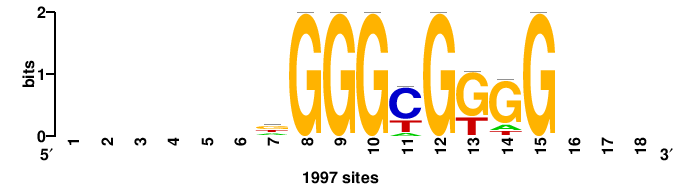
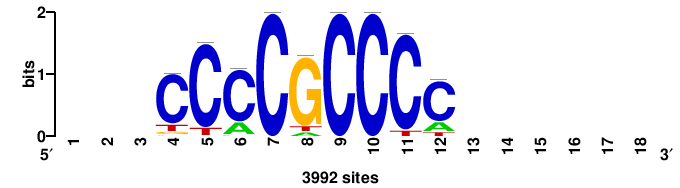
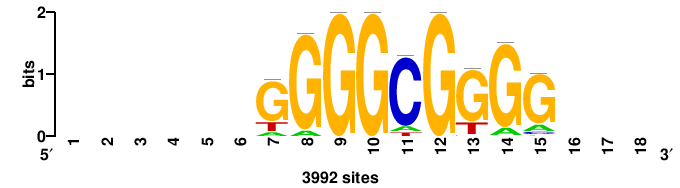
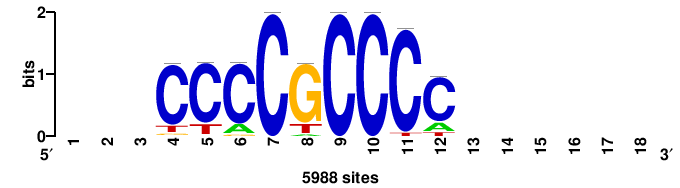
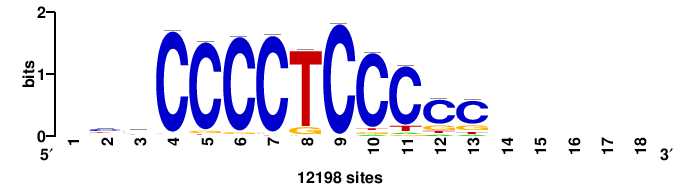
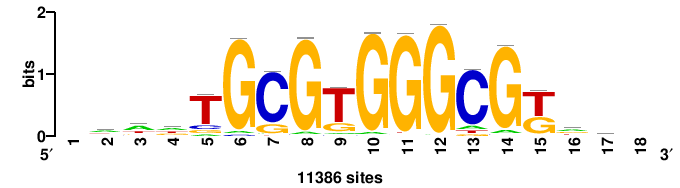
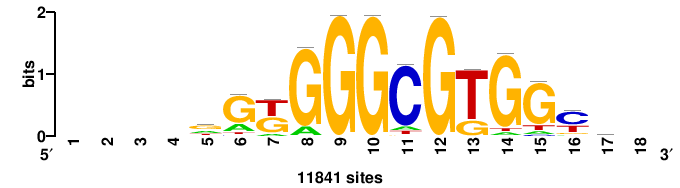
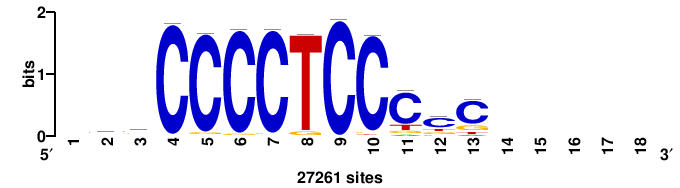
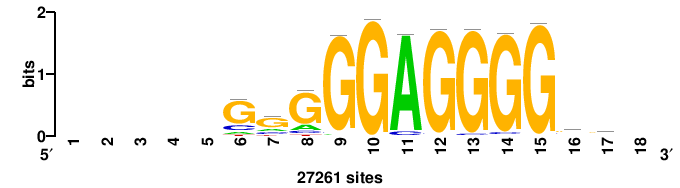
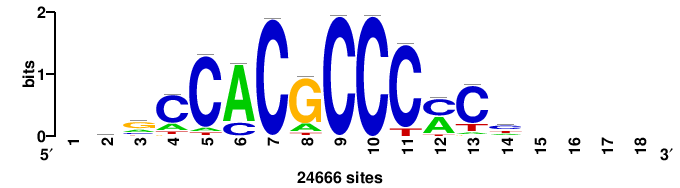
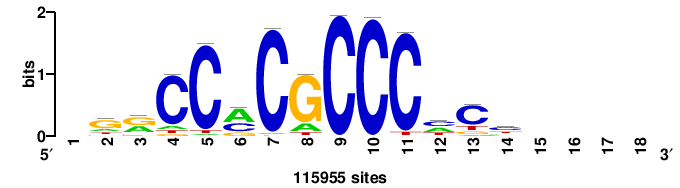
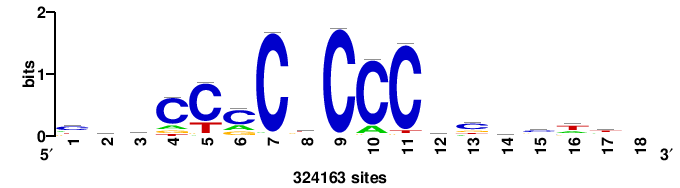
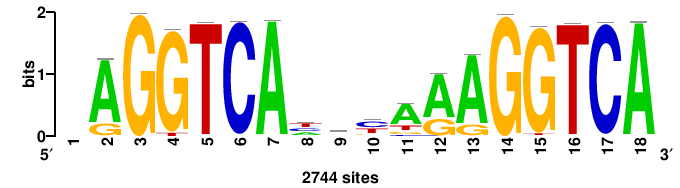
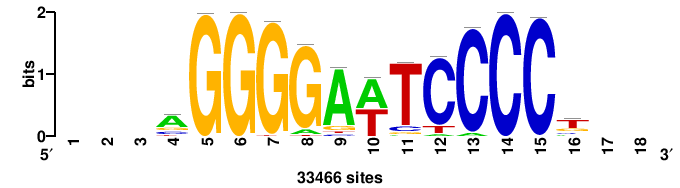
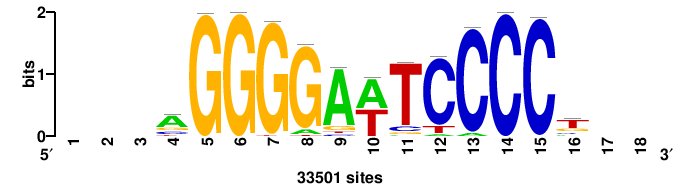
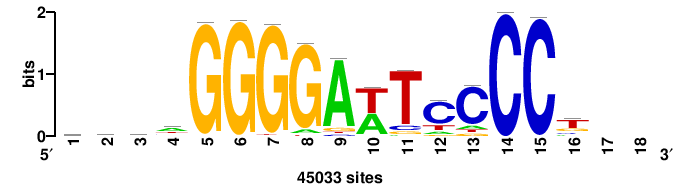
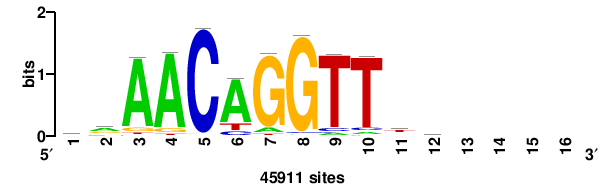
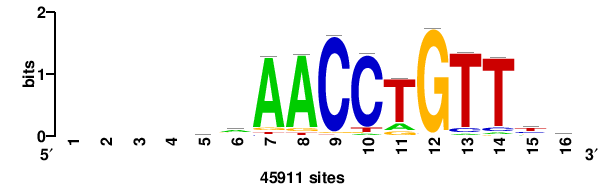
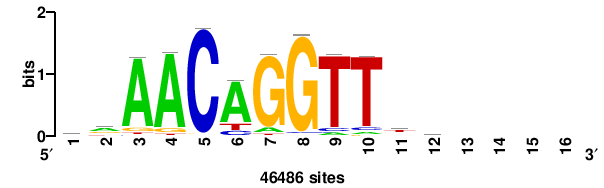
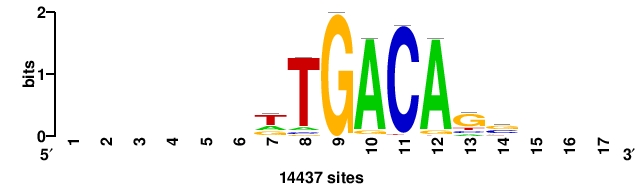
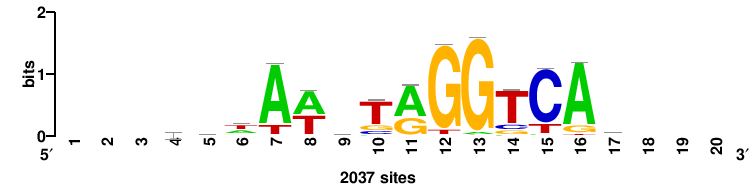
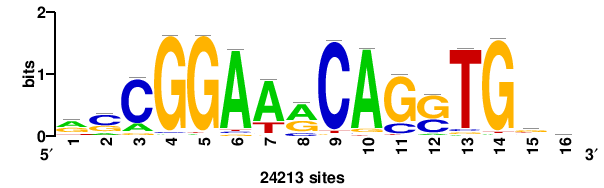
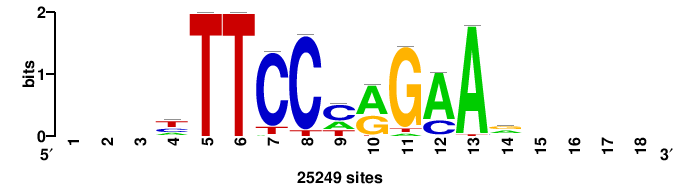
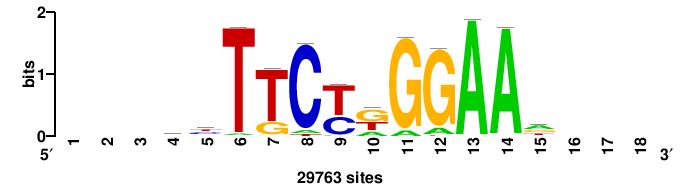
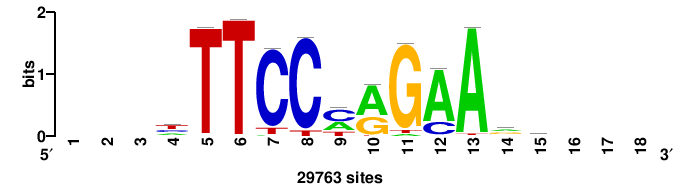
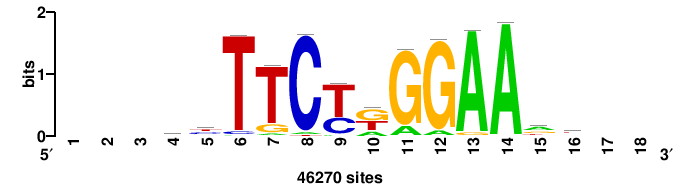
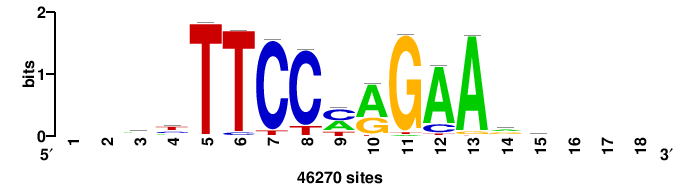
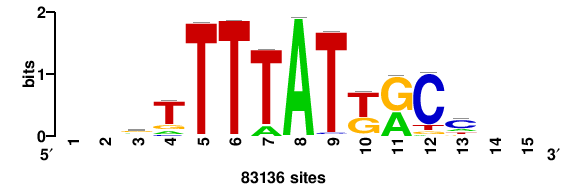
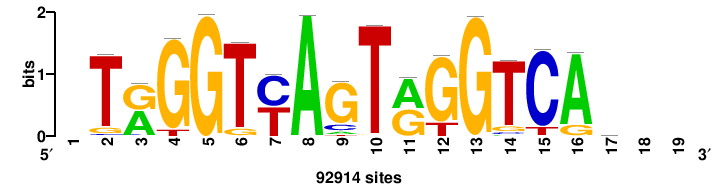
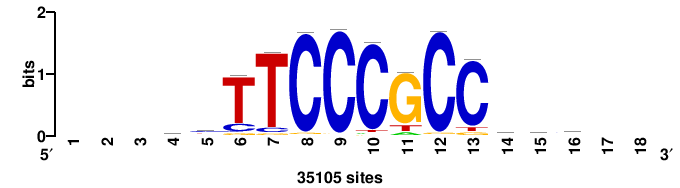
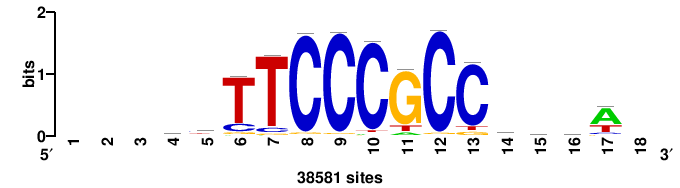
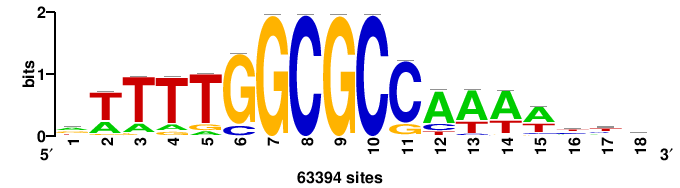
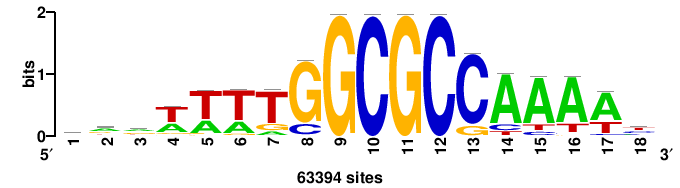
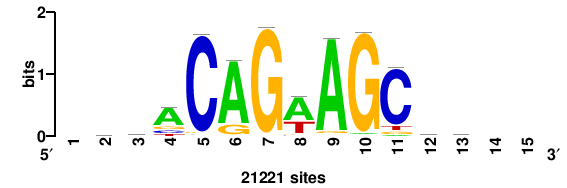
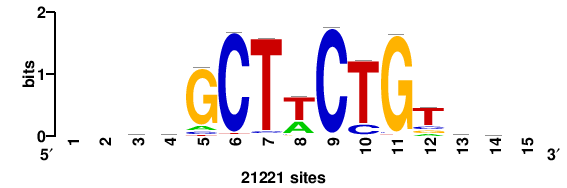
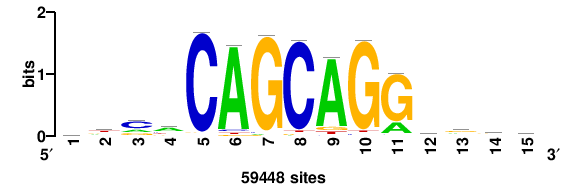
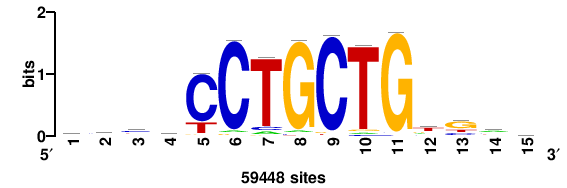
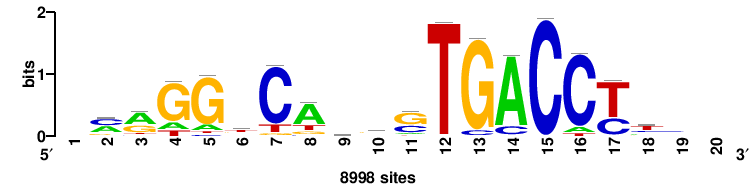
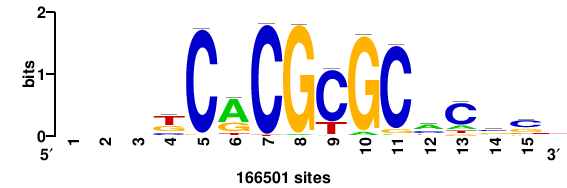
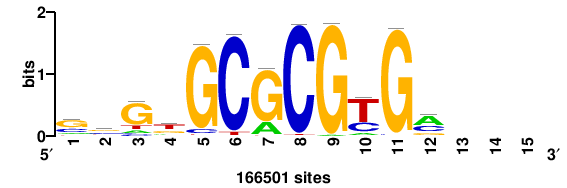
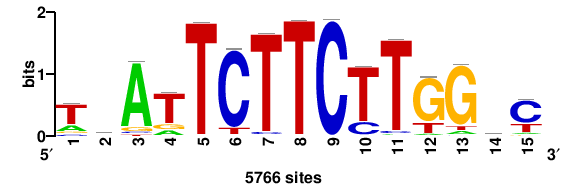
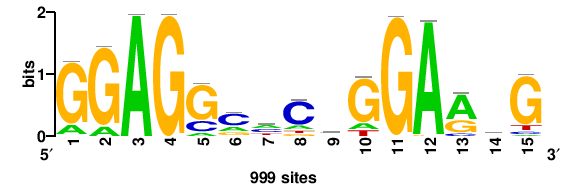
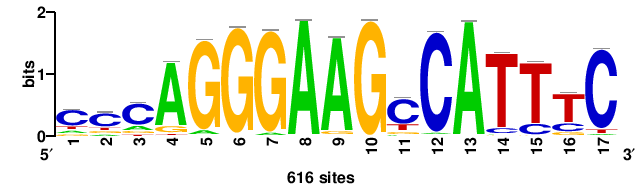
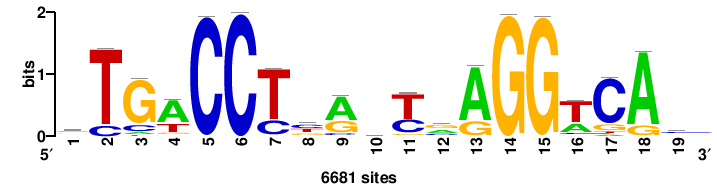
| JASPAR_2022_vertebrates_CORE_m328_MA0147_3 |
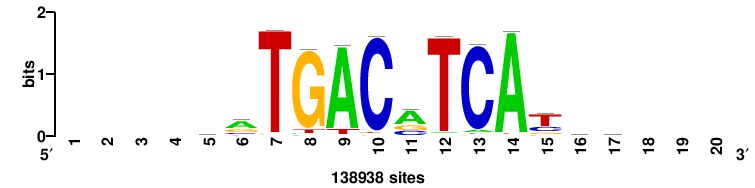
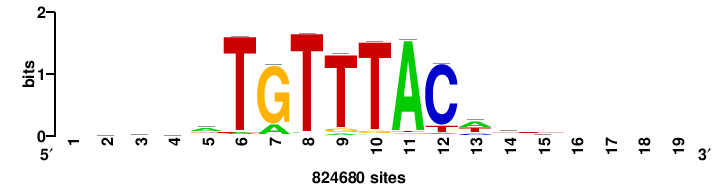
MYC |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.74 |
4942 |
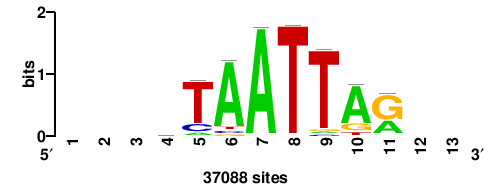
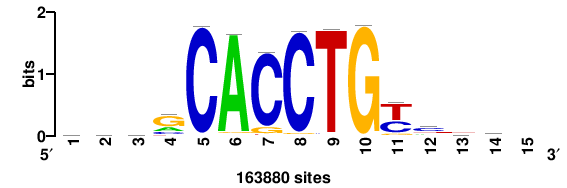
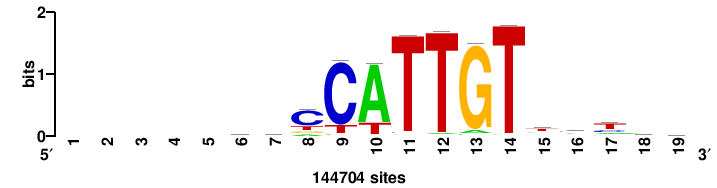
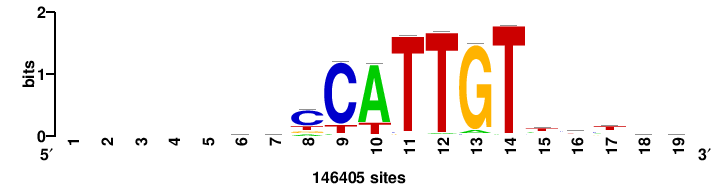
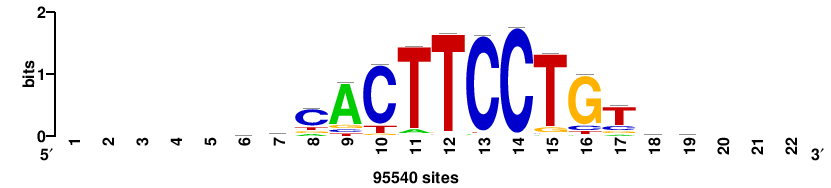
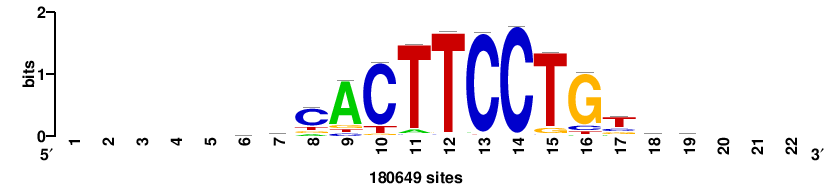
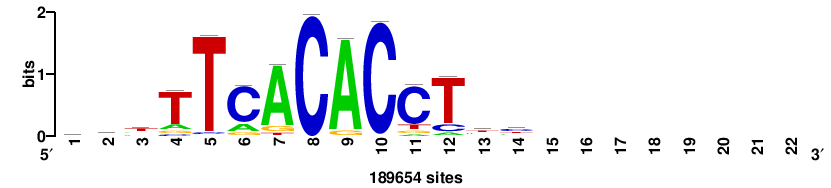
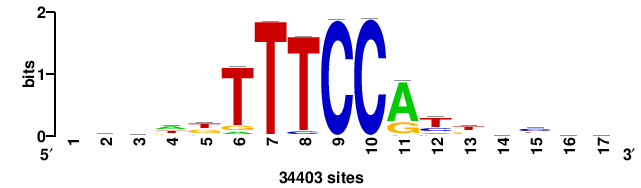
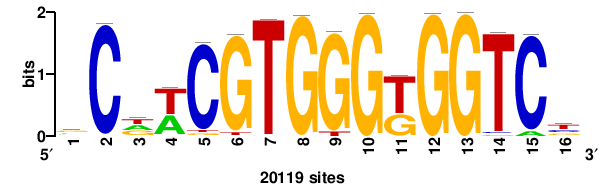
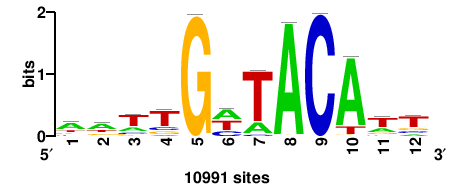
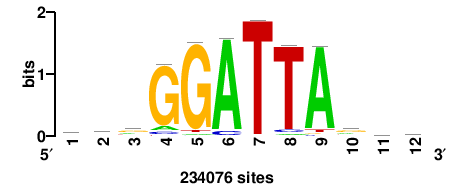
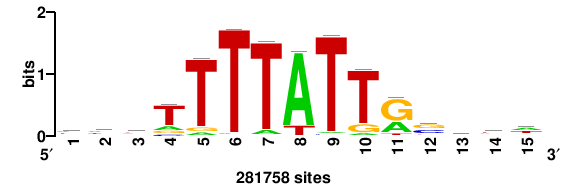
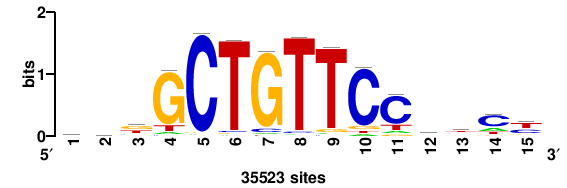
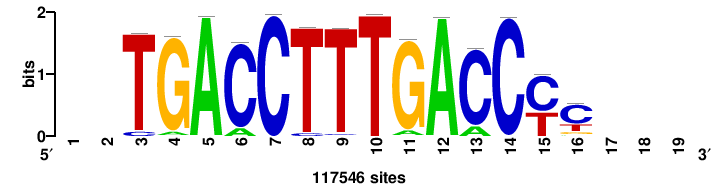
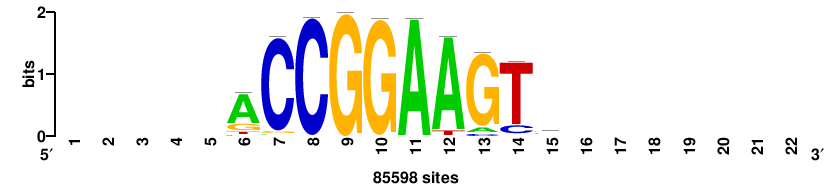
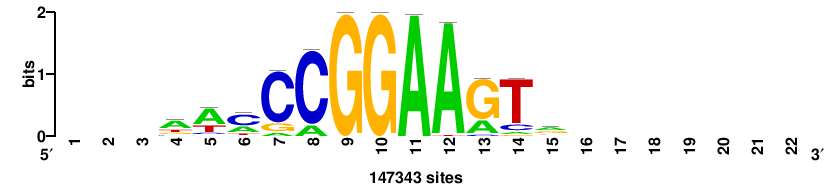
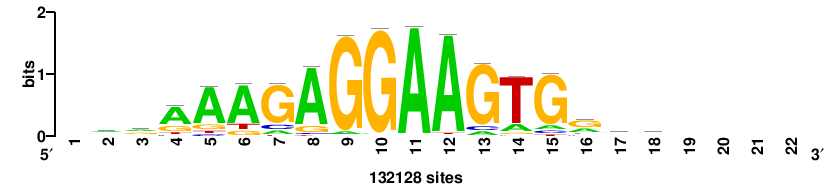
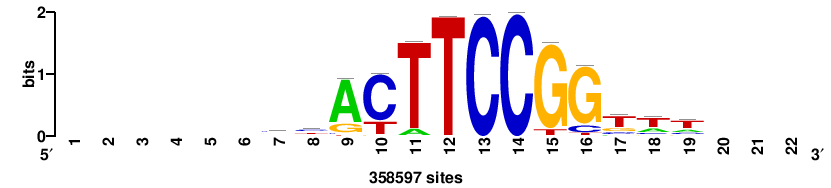
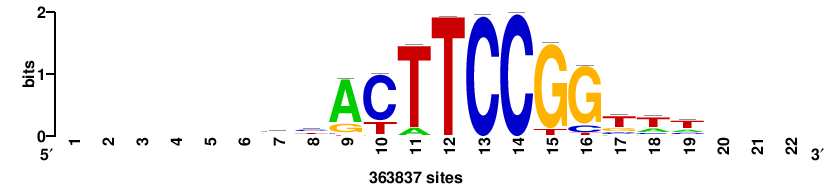
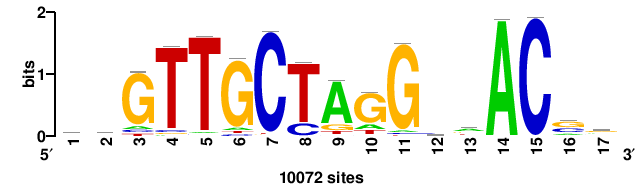
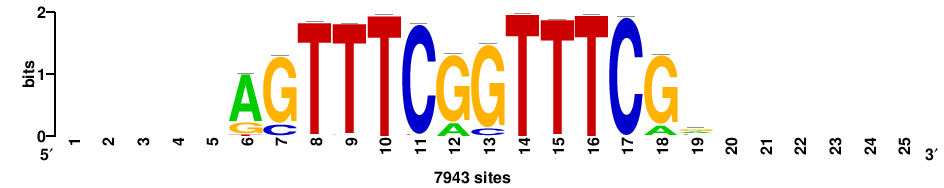
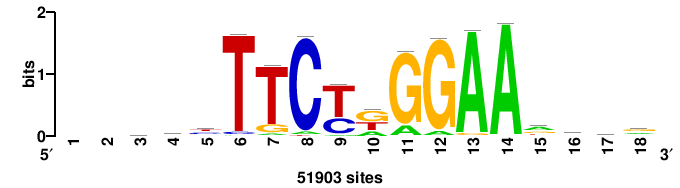
+++SRGCACGTGGSS+++ |
+++ssCCACGTGcys+++ |
 |
 |
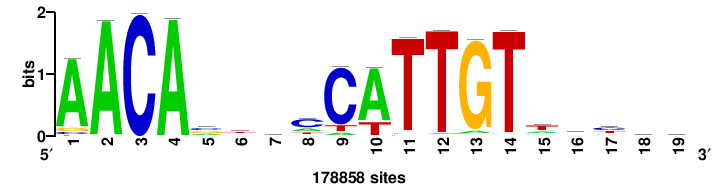
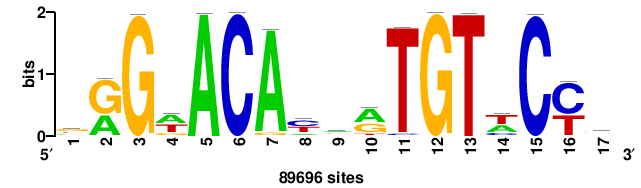
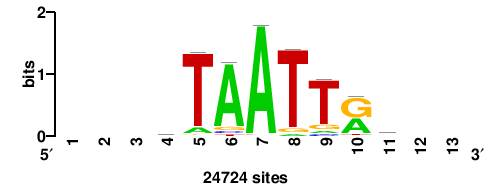
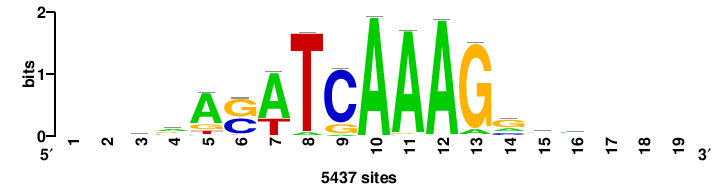
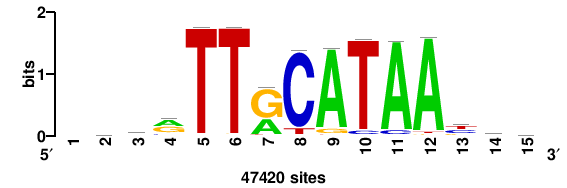
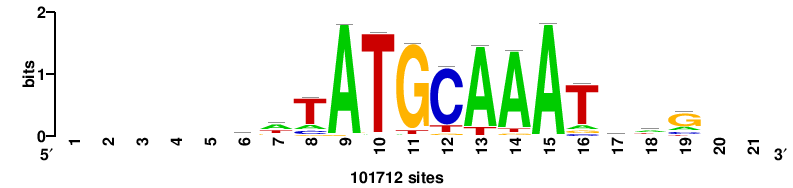
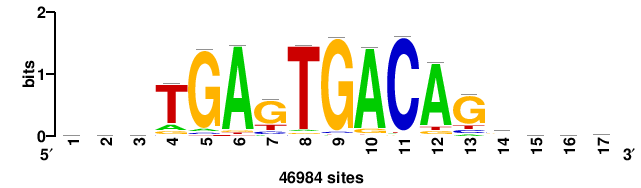
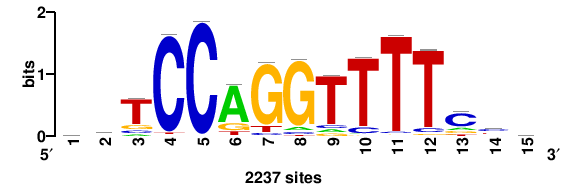
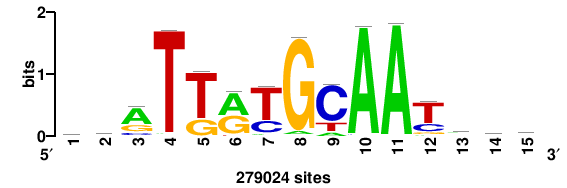
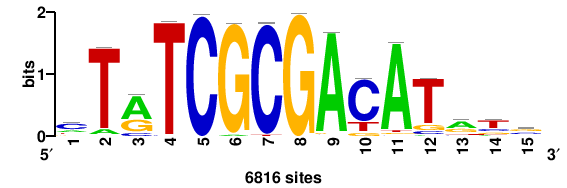
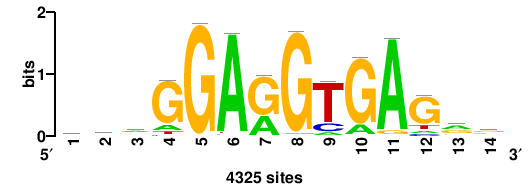
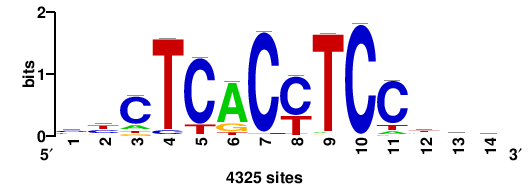
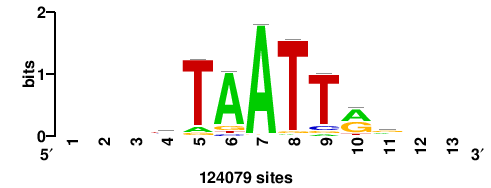
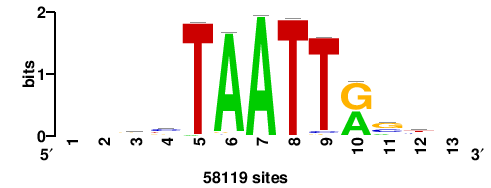
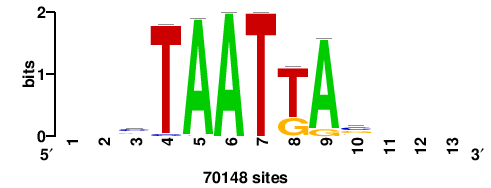
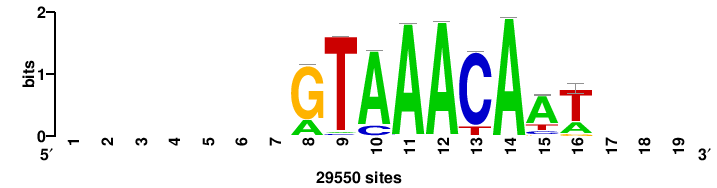
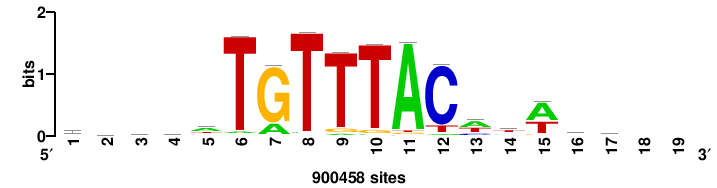
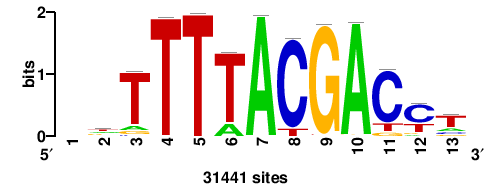
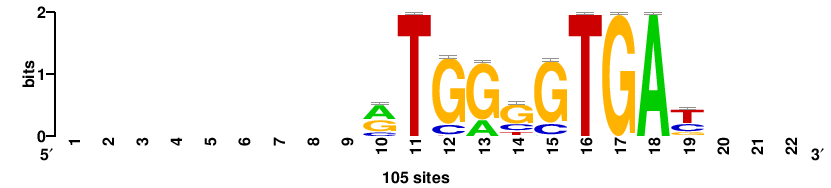
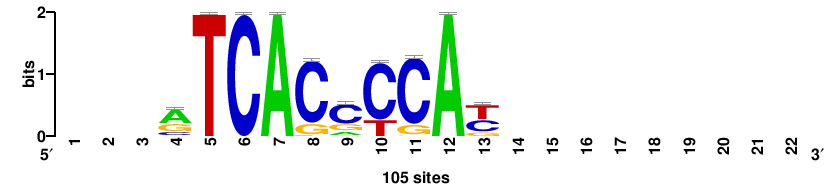
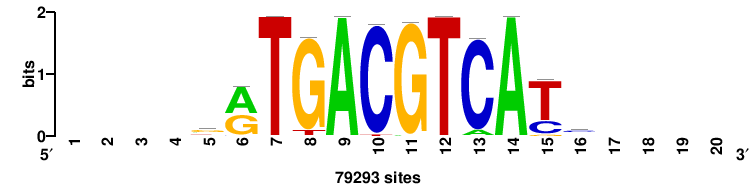
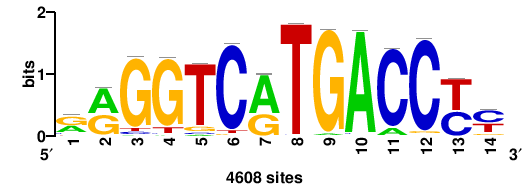
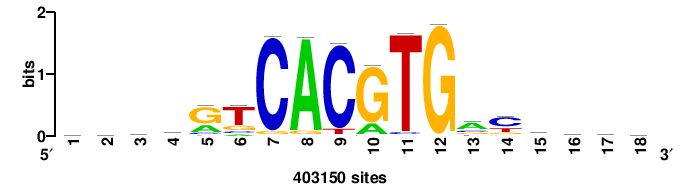
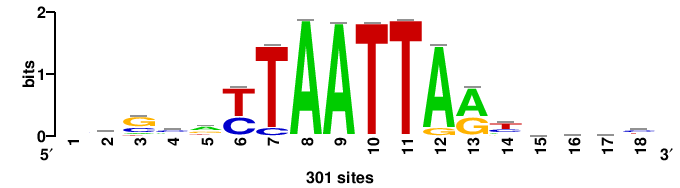
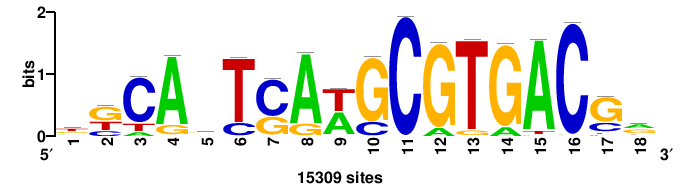
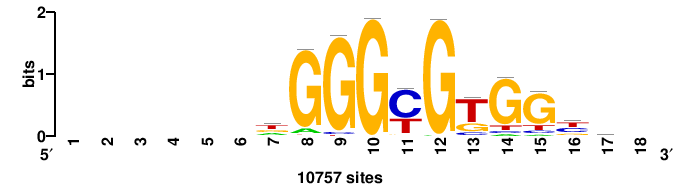
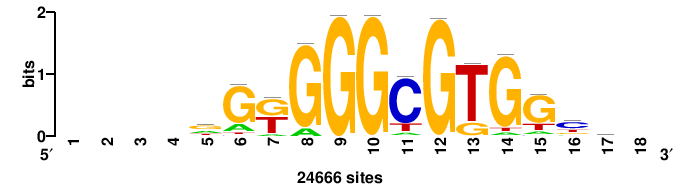
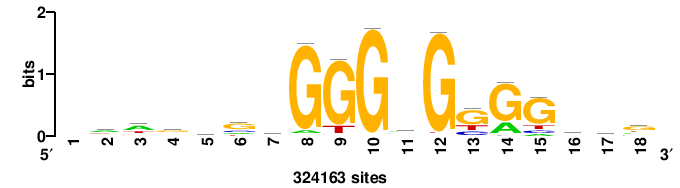
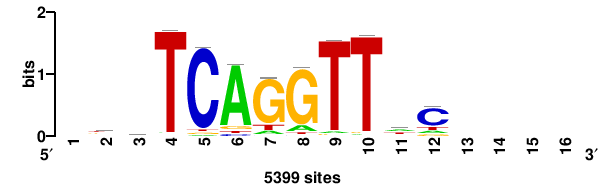
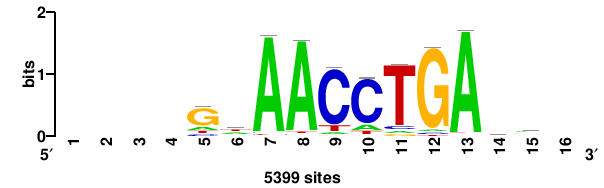
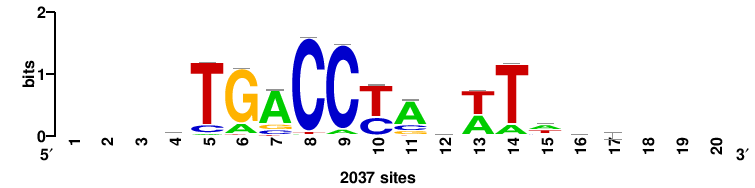
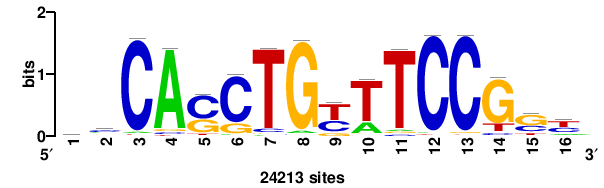
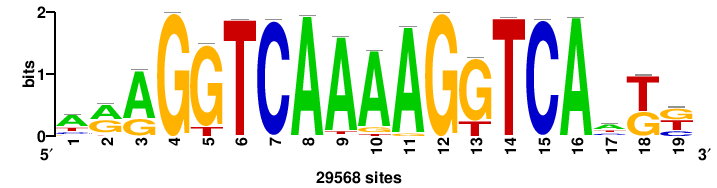
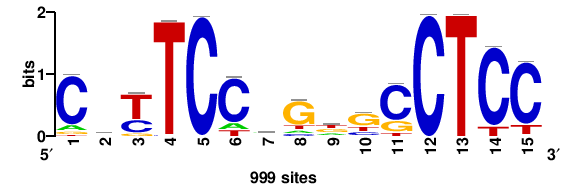
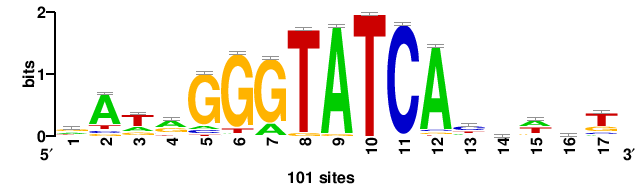
| JASPAR_2022_vertebrates_CORE_m809_MA0847_3 |
FOXD2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
9.93 |
4126 |
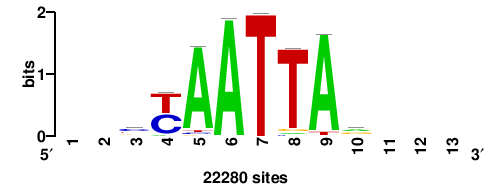
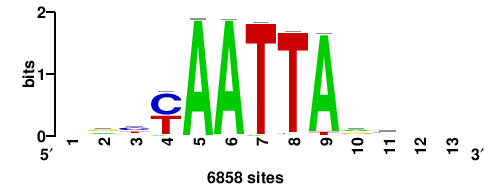
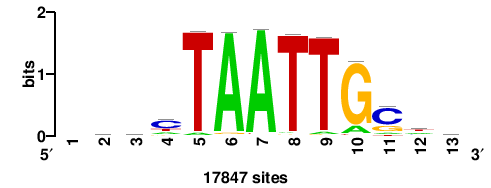
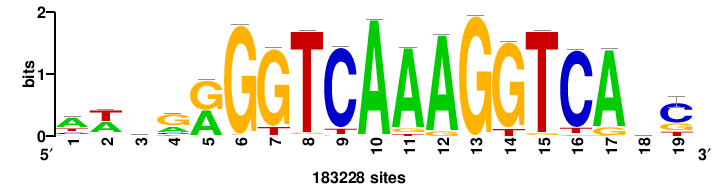
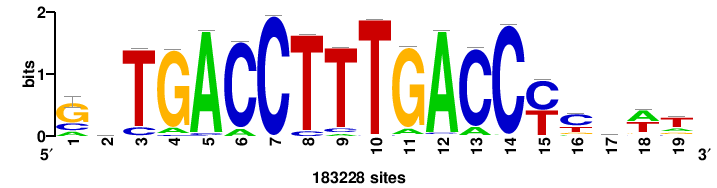
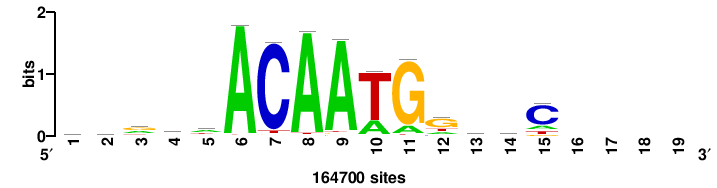
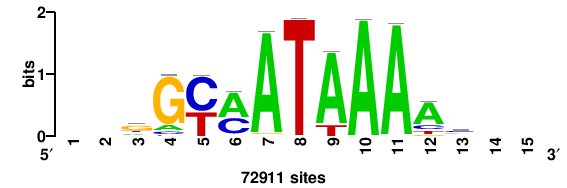
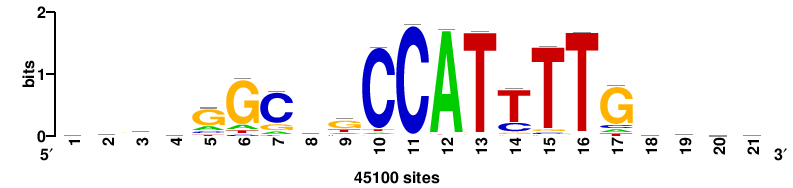
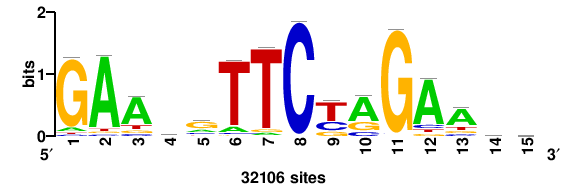
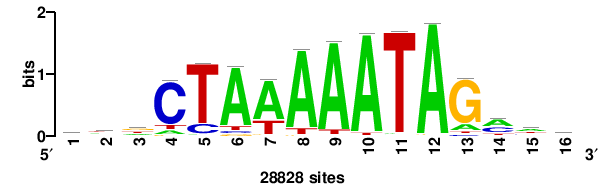
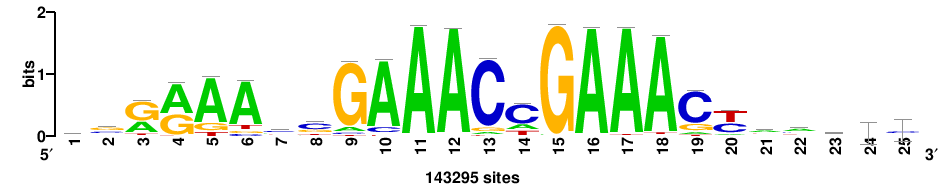
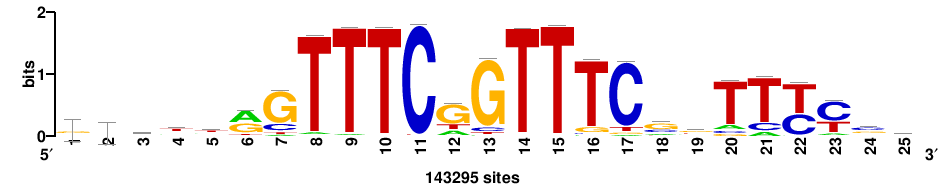
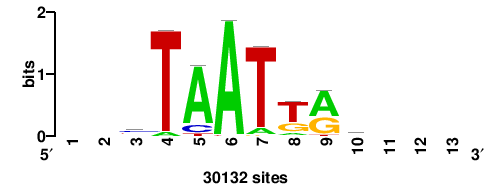
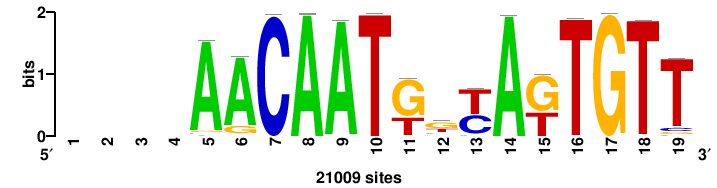
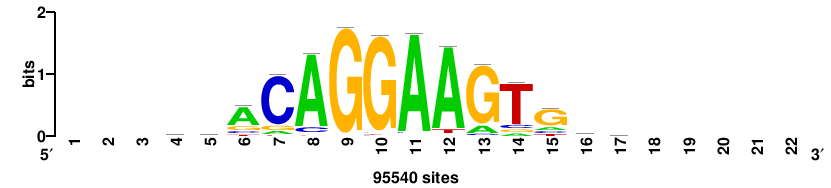
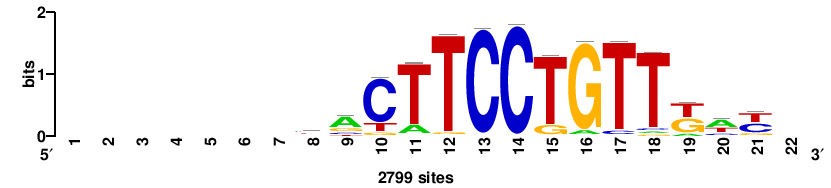
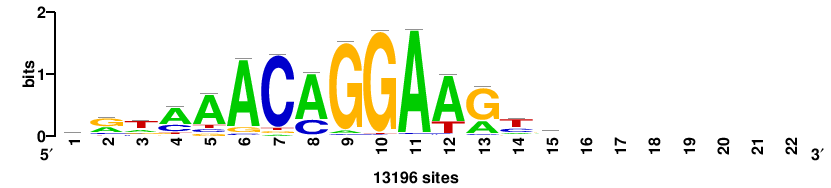
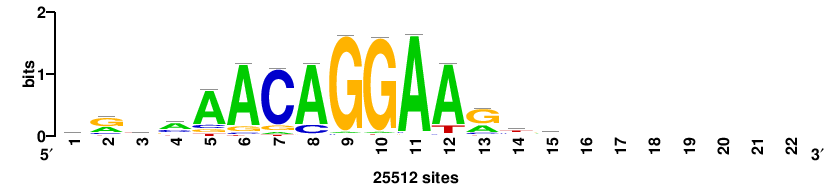
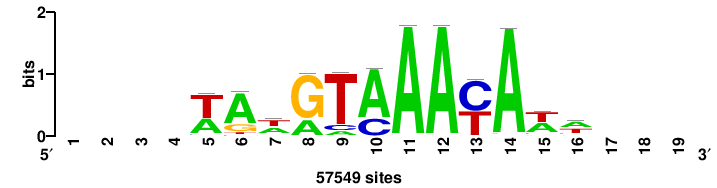
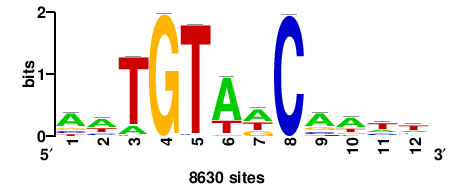
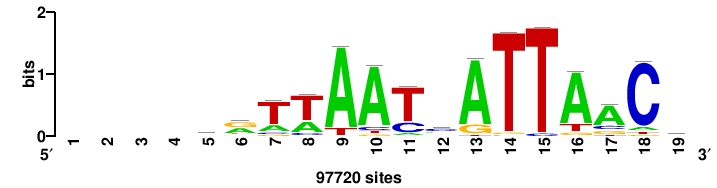
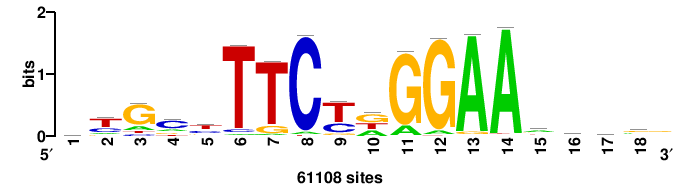
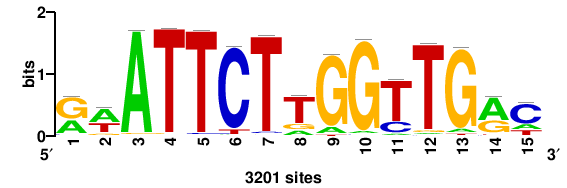
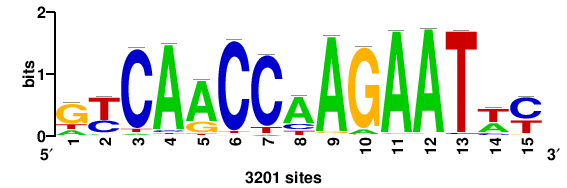
++syTAAryAAACA+++++ |
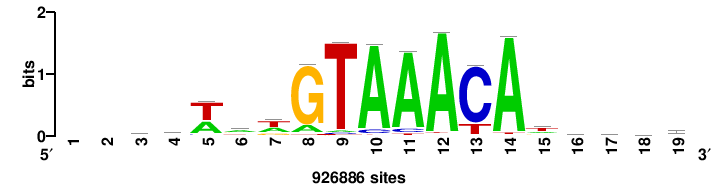
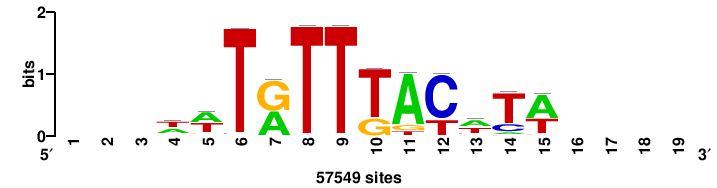
+++++TGTTTRYTTARS++ |
 |
 |
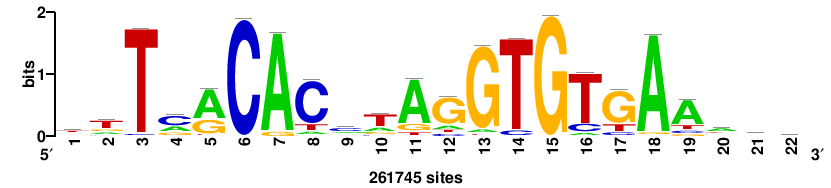
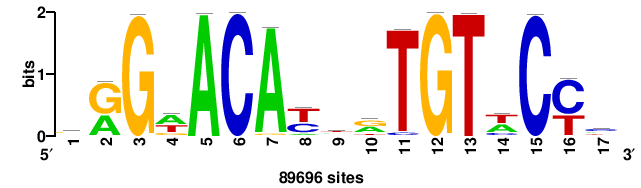
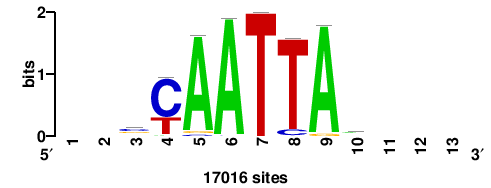
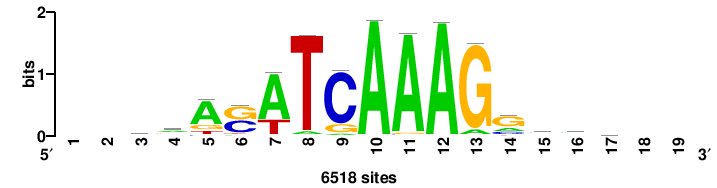
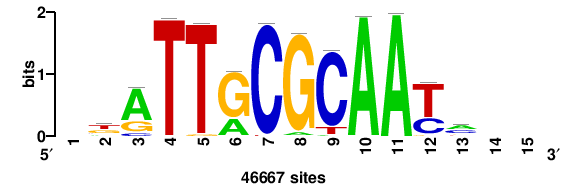
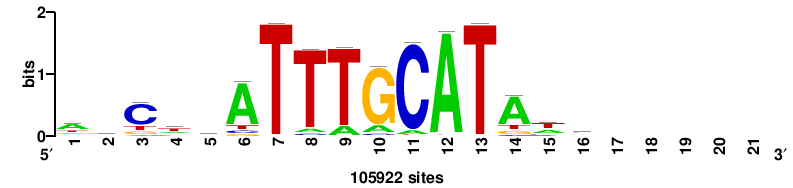
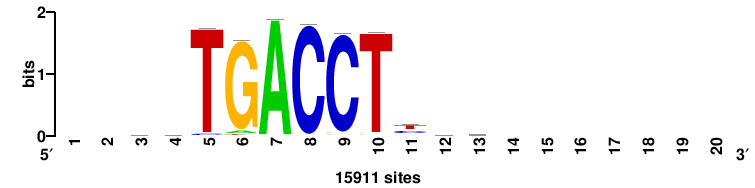
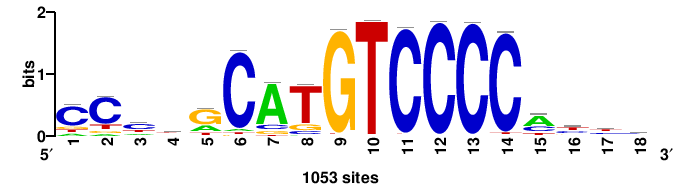
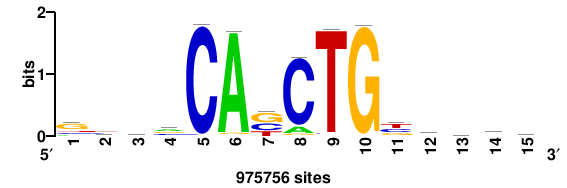
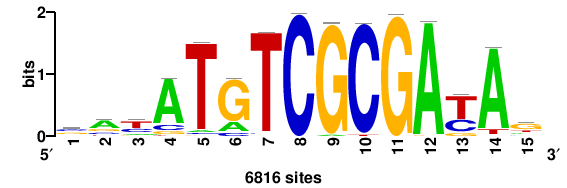
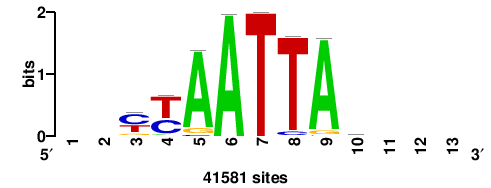
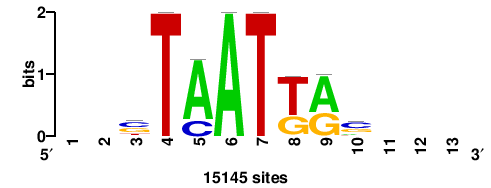
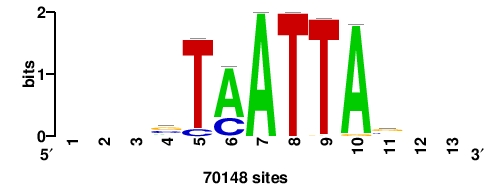
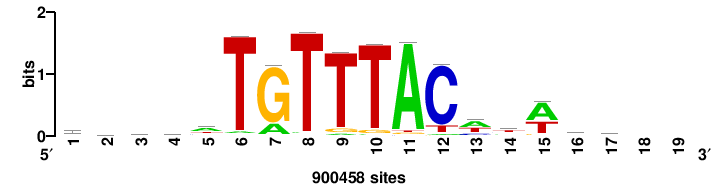
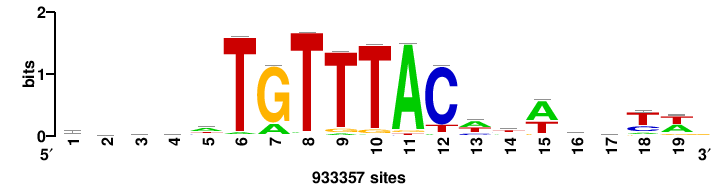
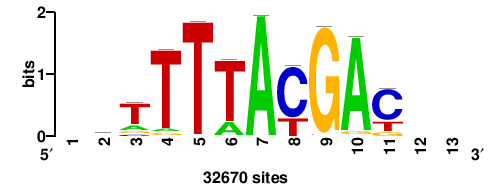
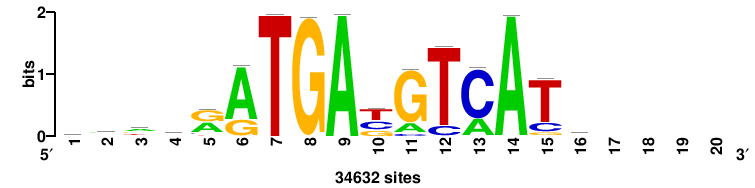
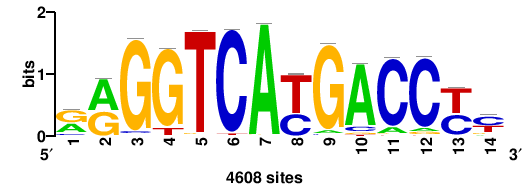
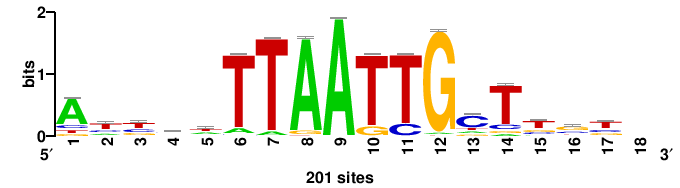
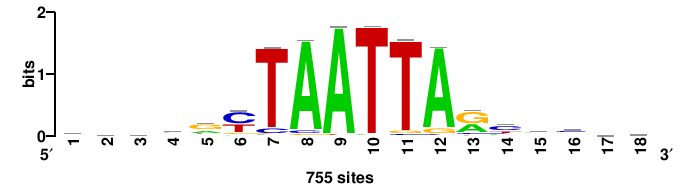
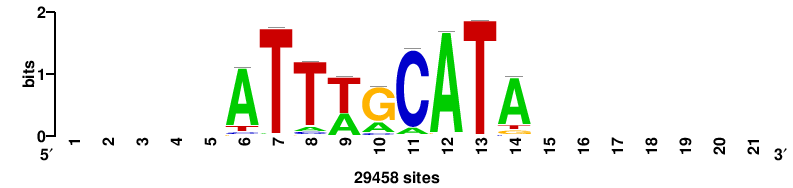
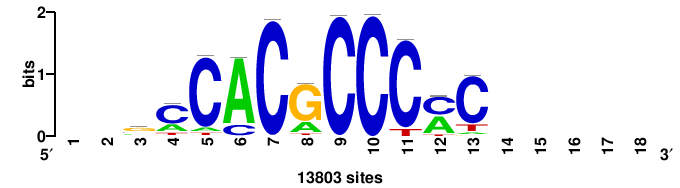
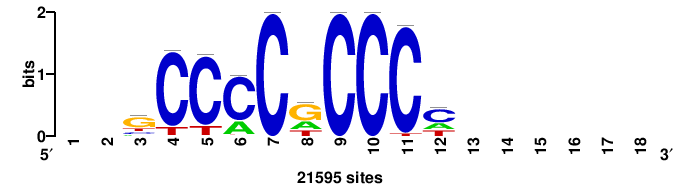
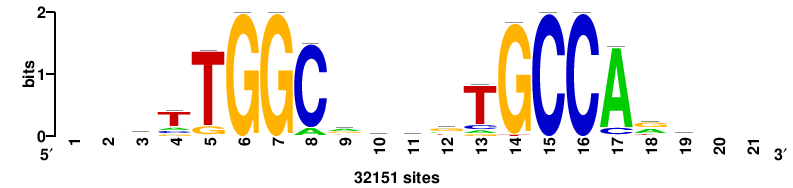
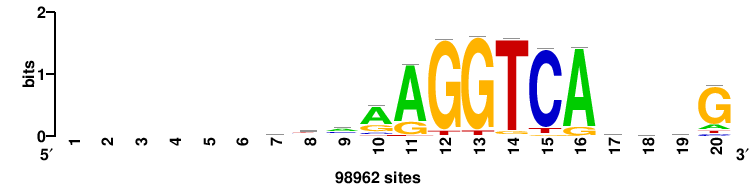
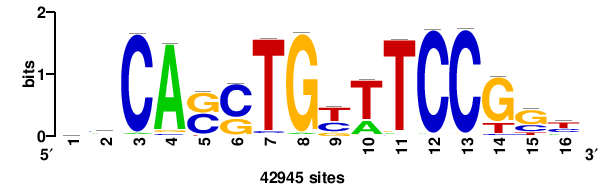
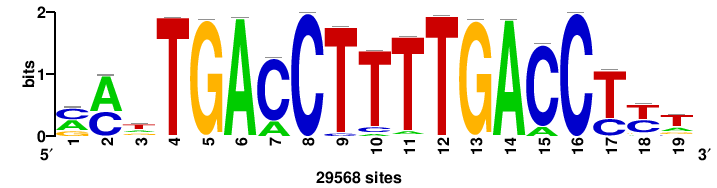
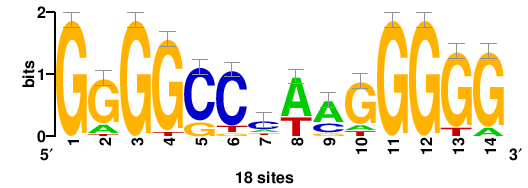
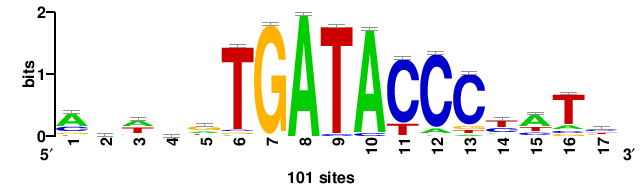
| JASPAR_2022_vertebrates_CORE_m743_MA0041_2 |
FOXD3 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.37 |
387 |
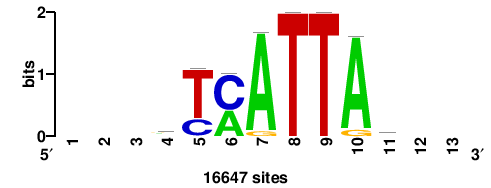
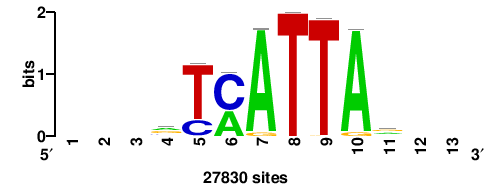
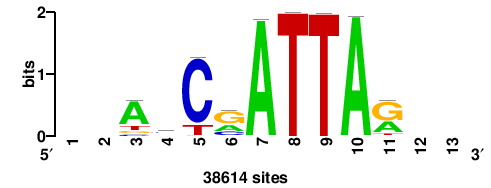
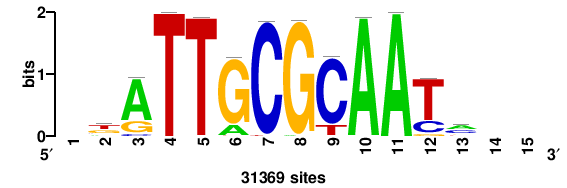
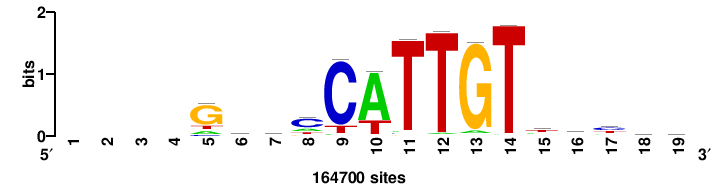
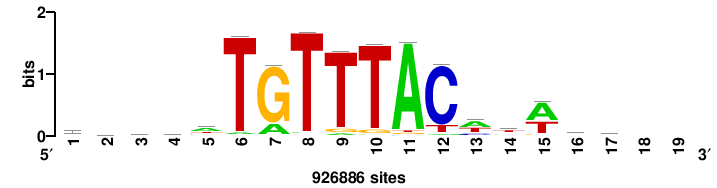
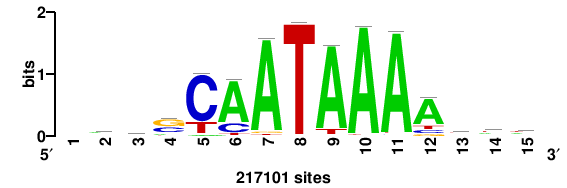
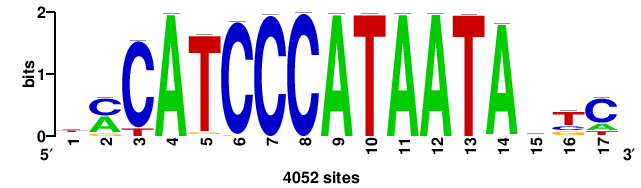
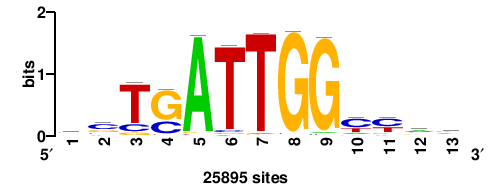
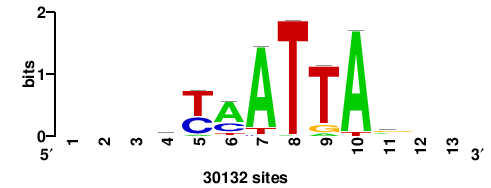
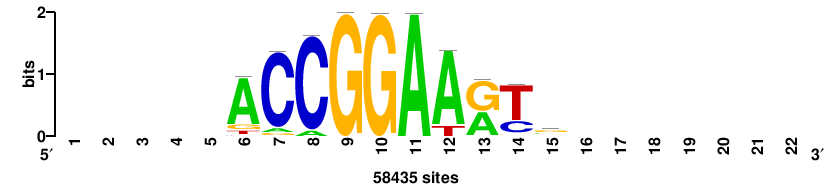
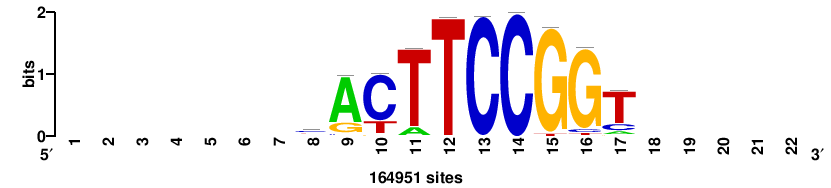
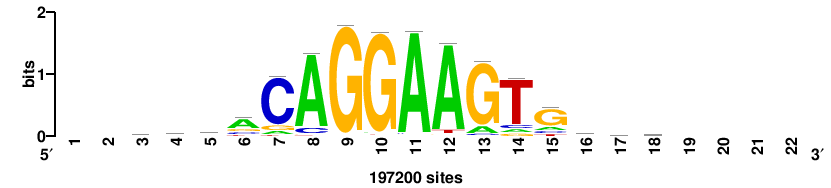
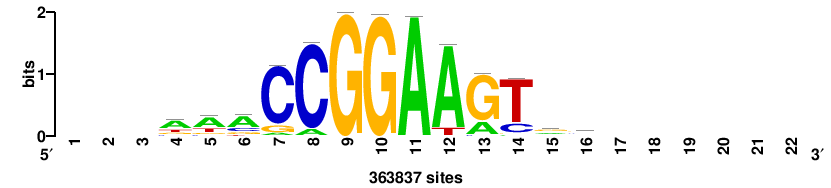
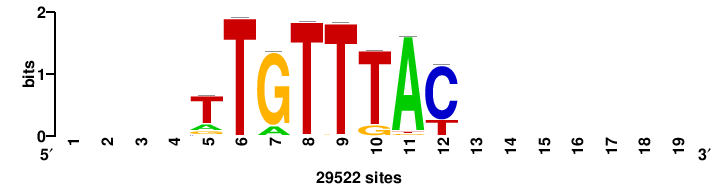
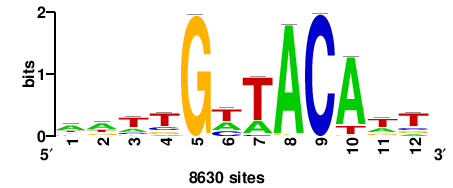
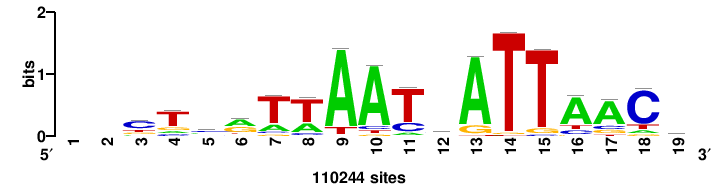
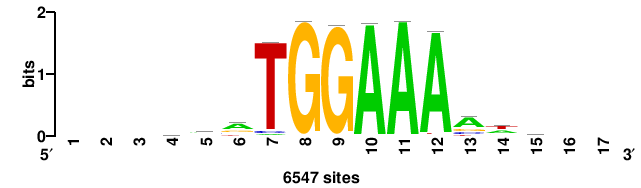
RTTGTTTRTTTDYWTW+++ |
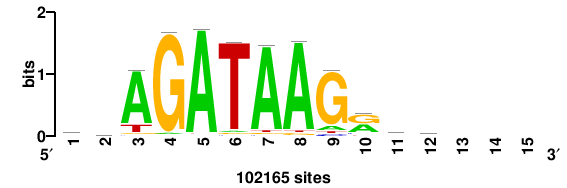
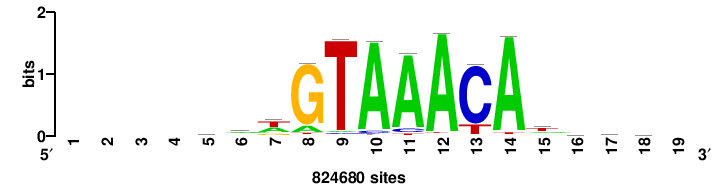
+++wawrhAAAYAAACaay |
 |
 |
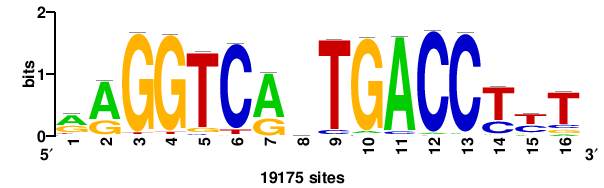
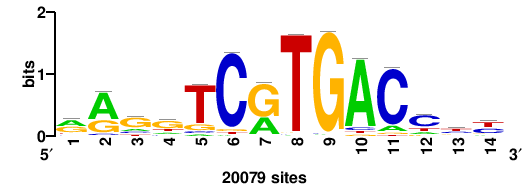
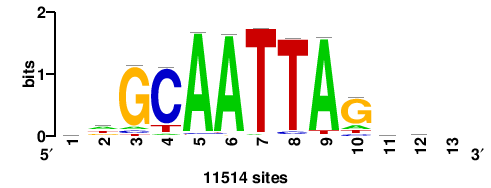
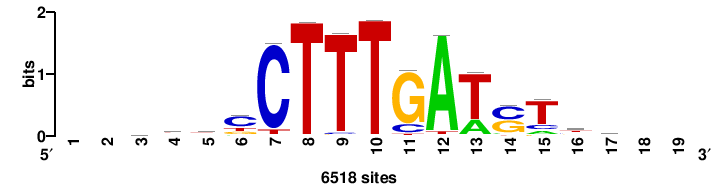
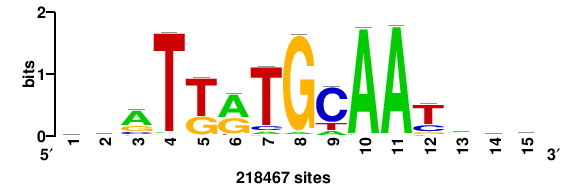
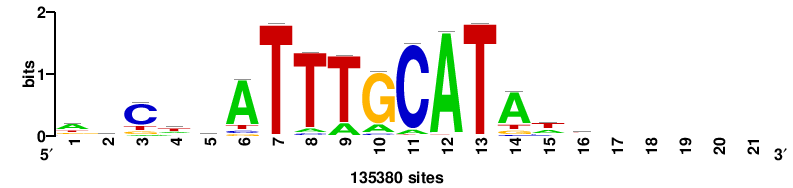
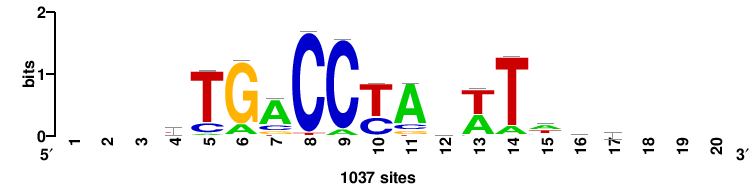
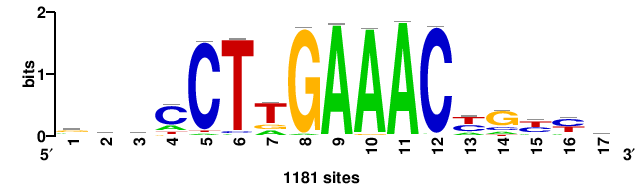
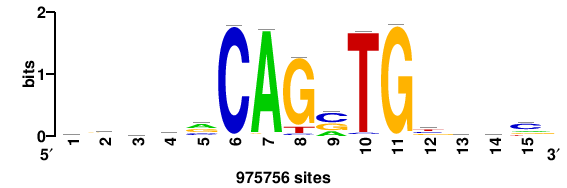
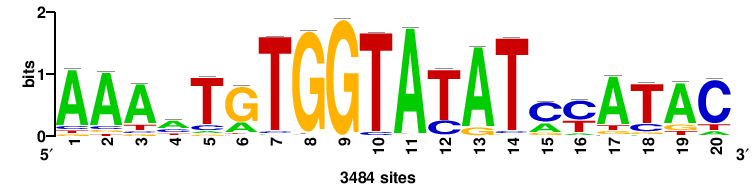
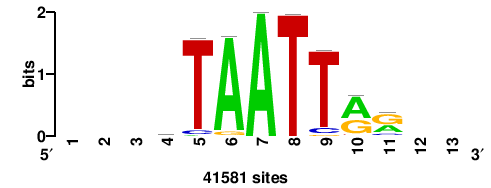
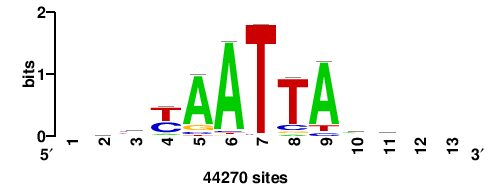
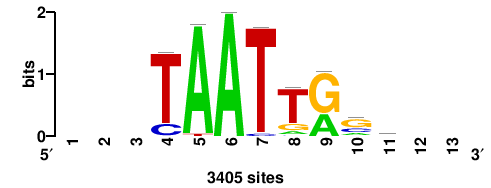
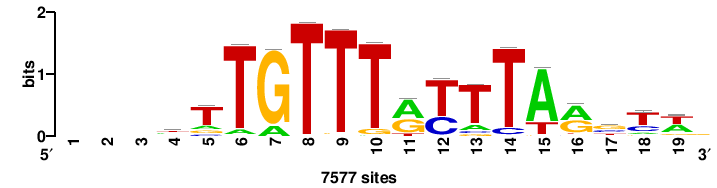
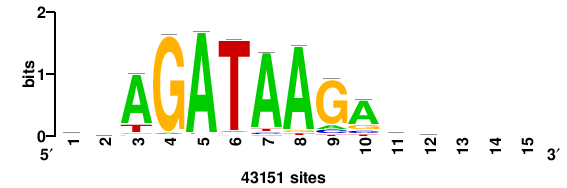
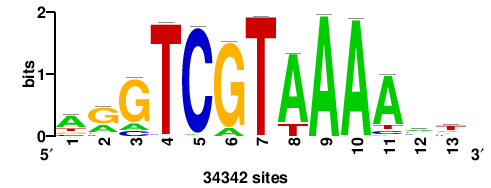
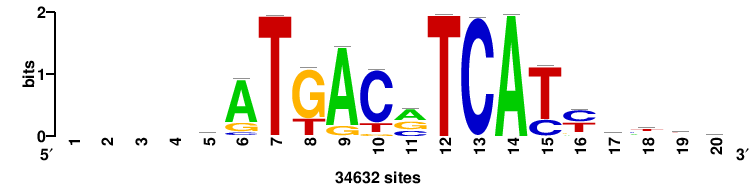
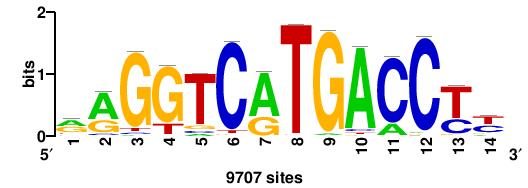
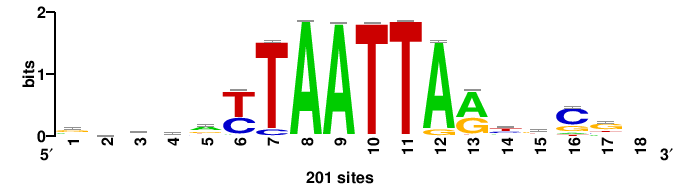
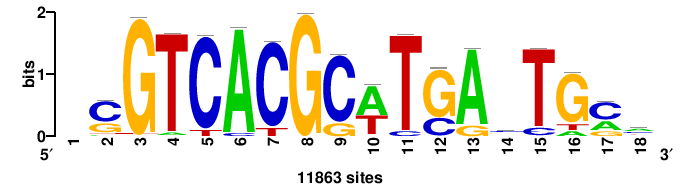
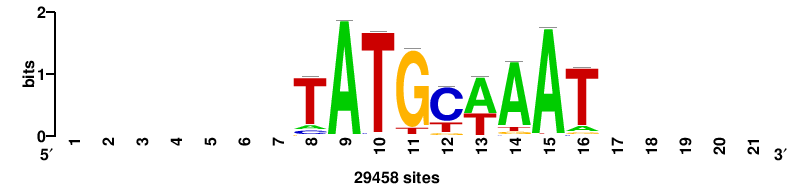
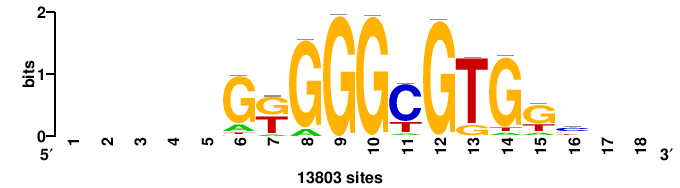
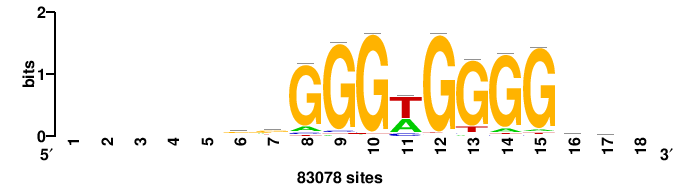
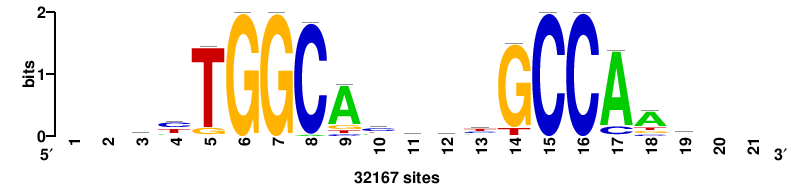
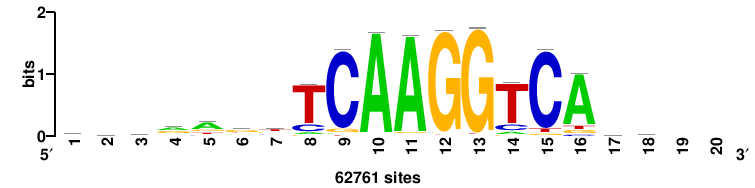
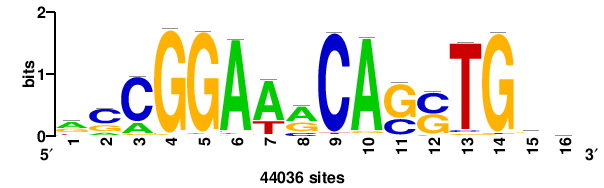
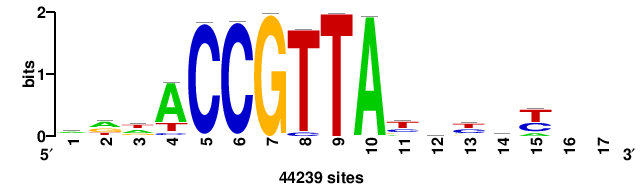
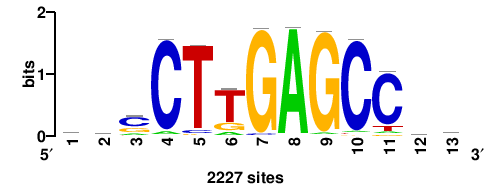
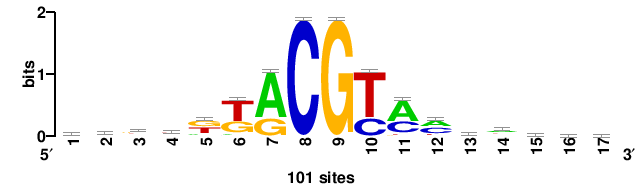
| JASPAR_2022_vertebrates_CORE_m783_MA0680_2 |
Pax7 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
9.22 |
41429 |
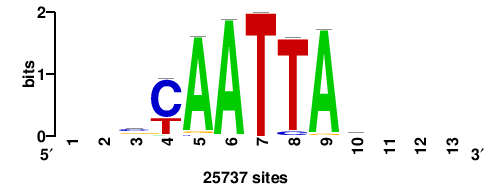
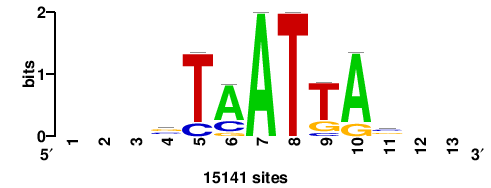
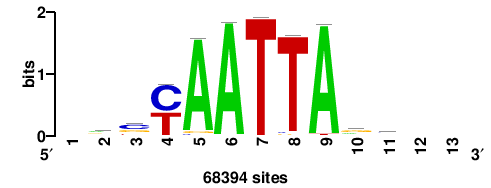
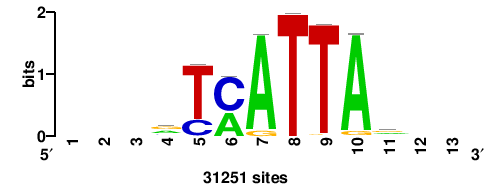
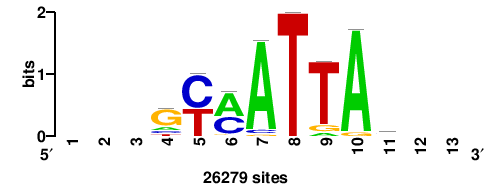
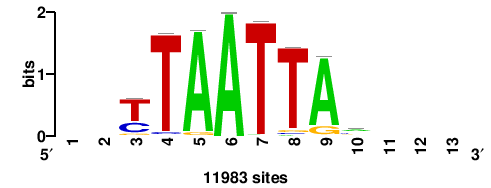
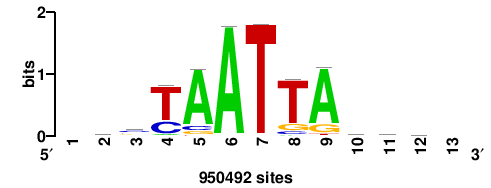
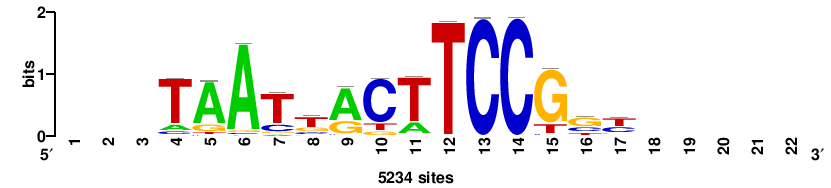
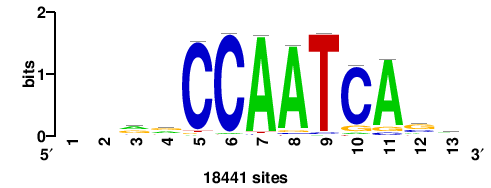
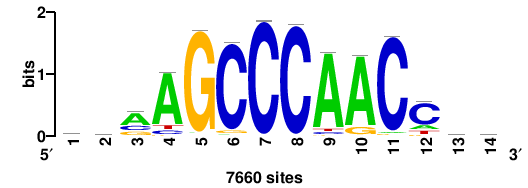
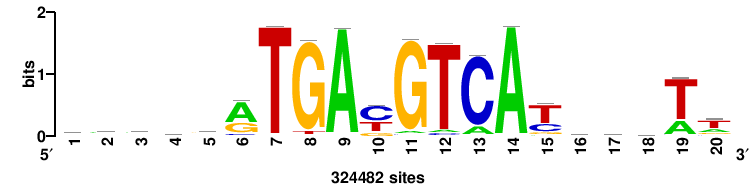
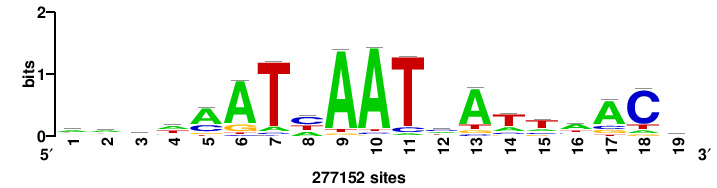
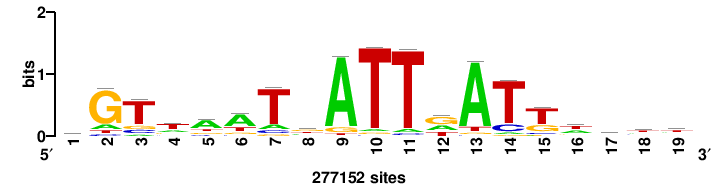
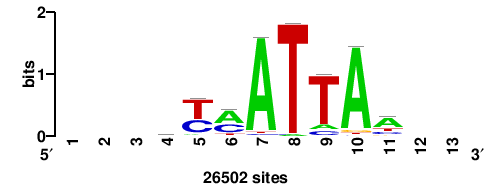
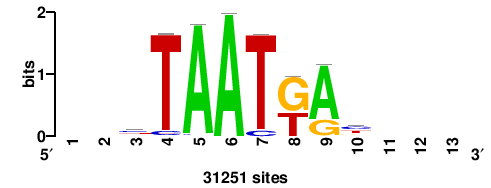
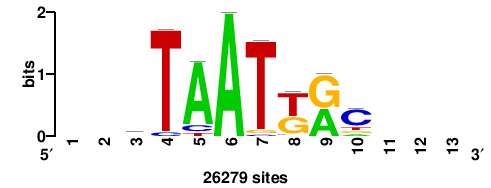
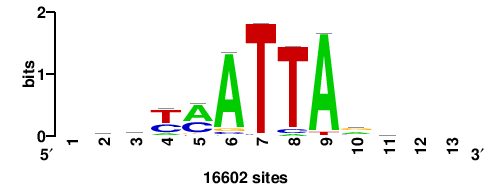
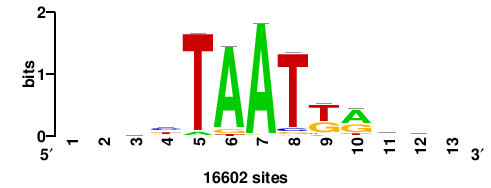
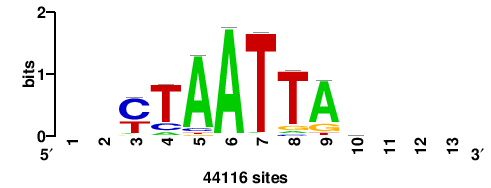
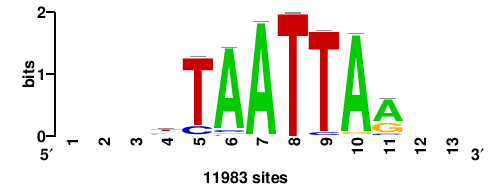
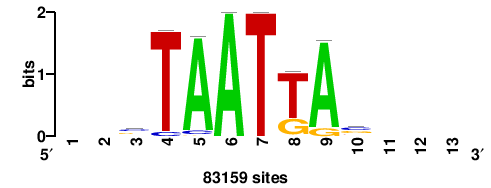
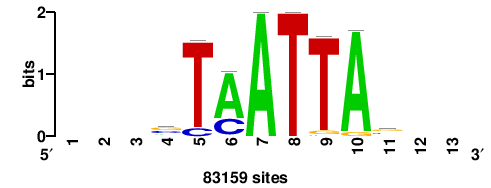
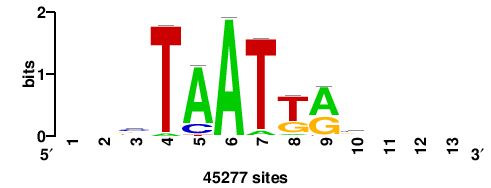
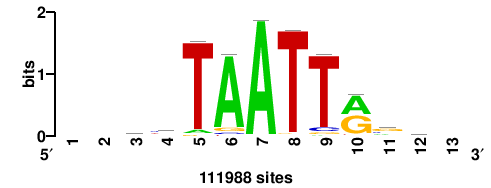
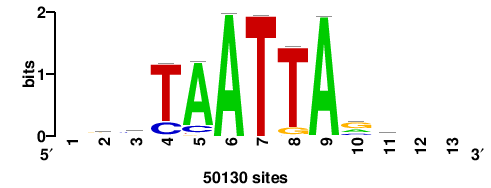
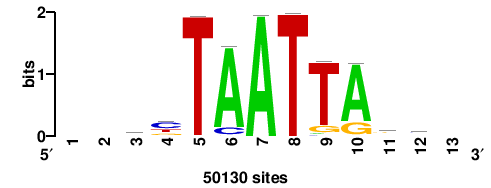
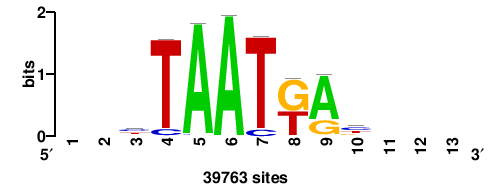
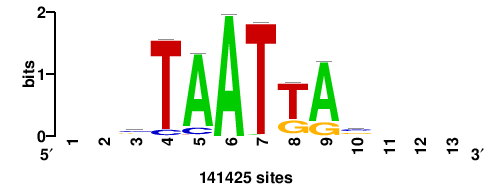
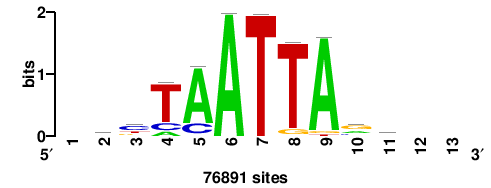
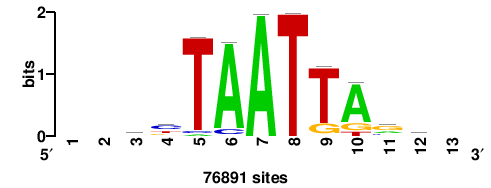
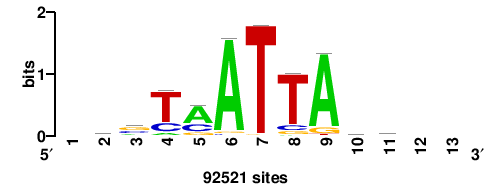
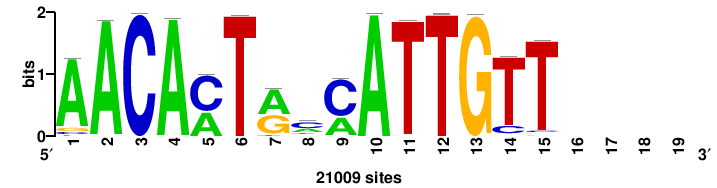
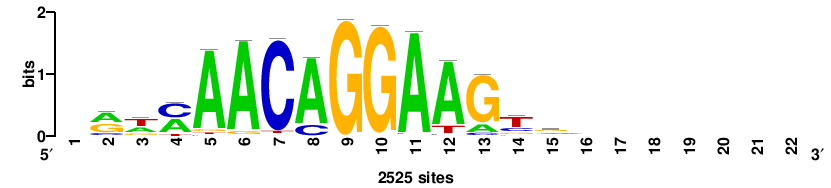
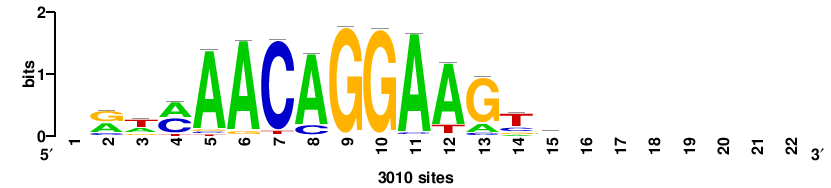
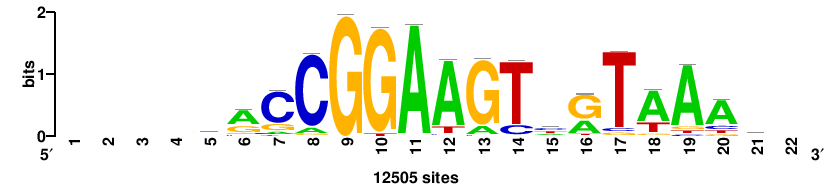
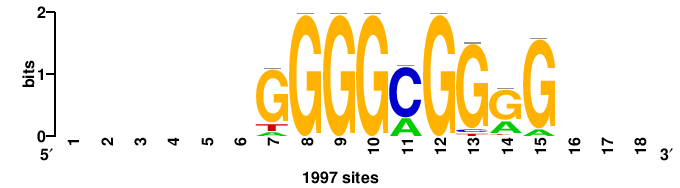
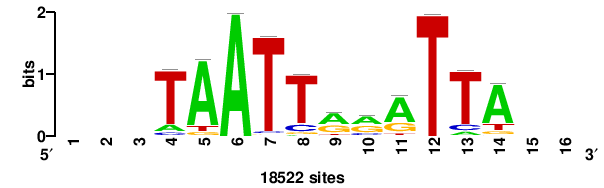
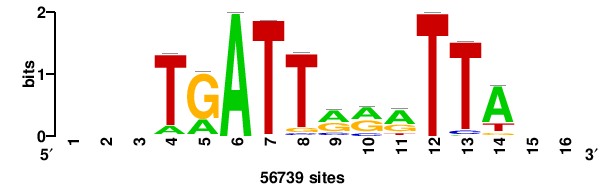
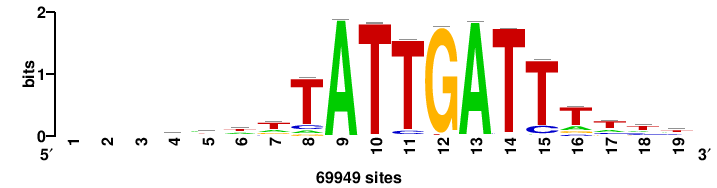
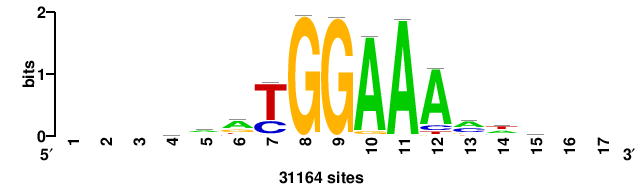
+wwtAATCAATTAwww+++ |
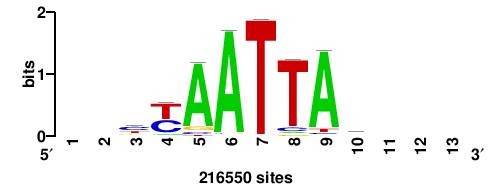
+++WWWTAATTGATTAWW+ |
 |
 |
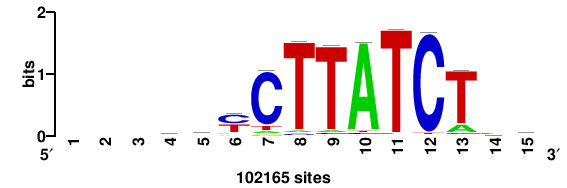
| JASPAR_2022_vertebrates_CORE_m271_MA0844_1 |
XBP1 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
10.73 |
12830 |
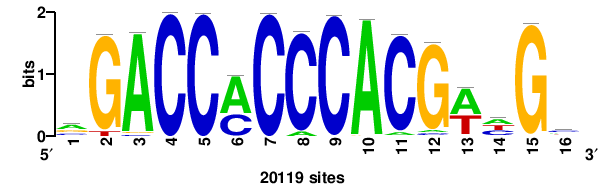
+++GMTGACGTGKCMYW+ |
+wrkGmCACGTCAkc+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m286_MA0860_1 |
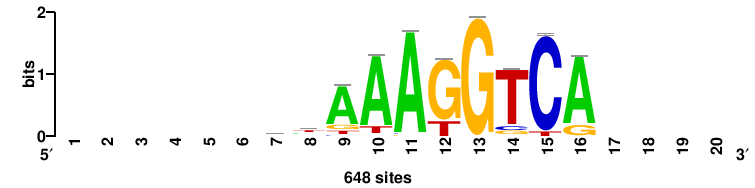
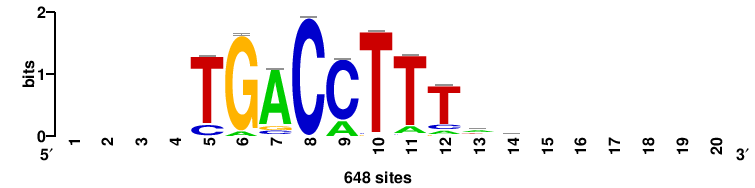
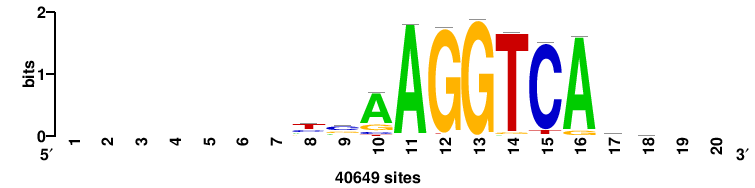
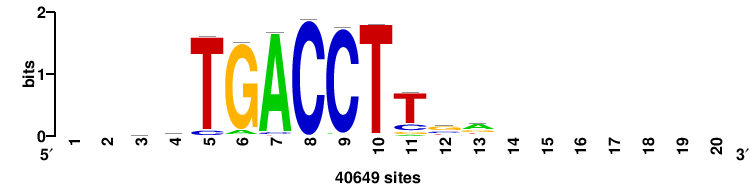
Rarg |
cluster_58 |
JASPAR_2022_vertebrates_CORE |
24 |
17.69 |
6258 |
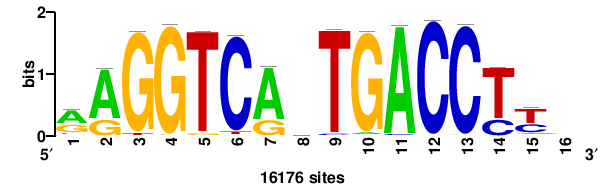
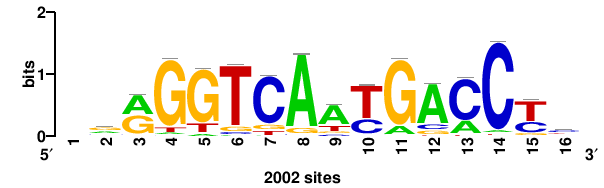
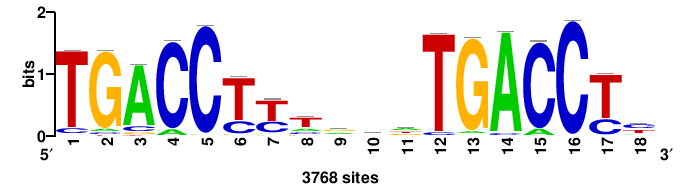
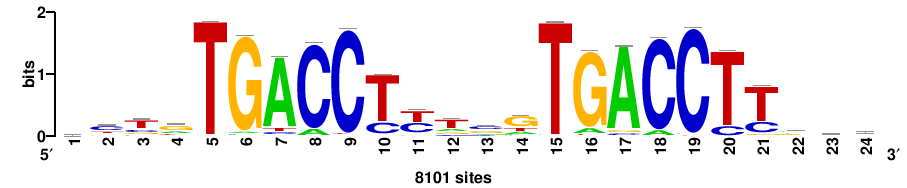
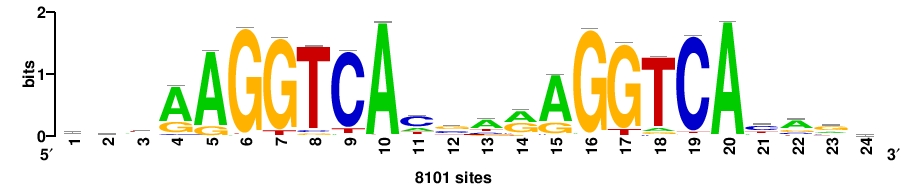
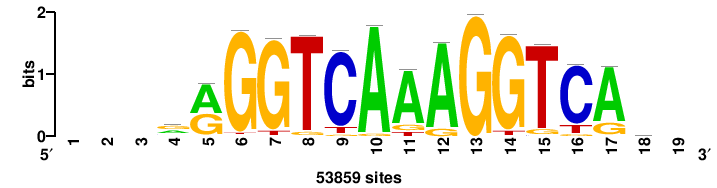
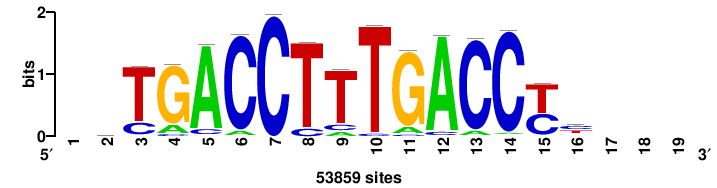
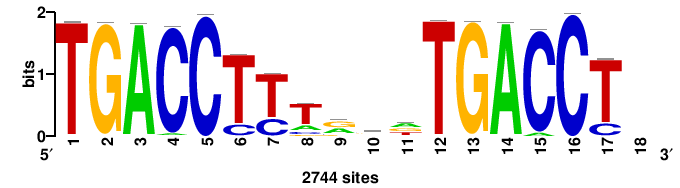
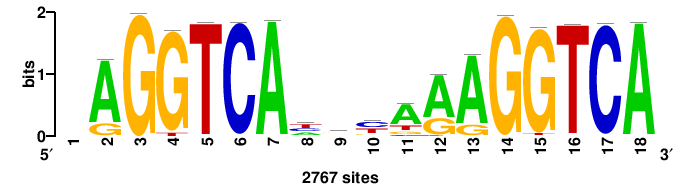
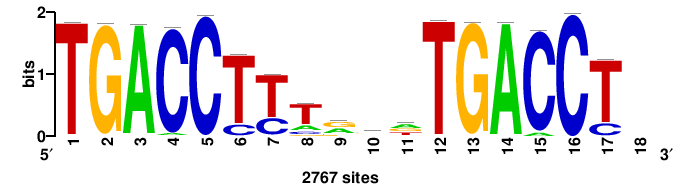
++++TGACCTYTSKTGACCTT+++ |
+++AAGGTCAmsArAGGTCA++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m461_MA1568_1 |
TCF21 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.25 |
1438 |
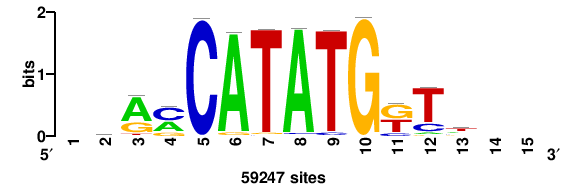
+caCCATATGkyr++ |
++YRMCATATGGTG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m185_MA0732_1 |
EGR3 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
12.58 |
2292 |
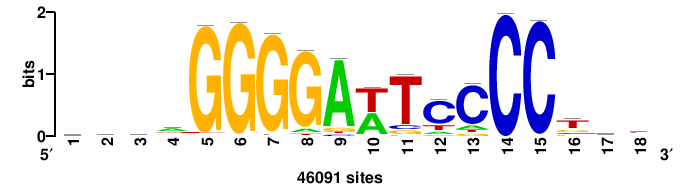
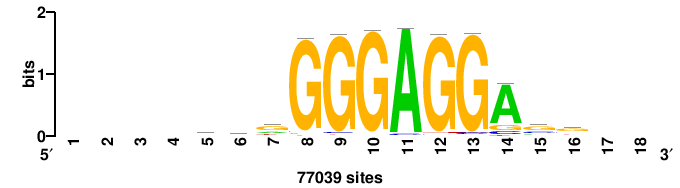
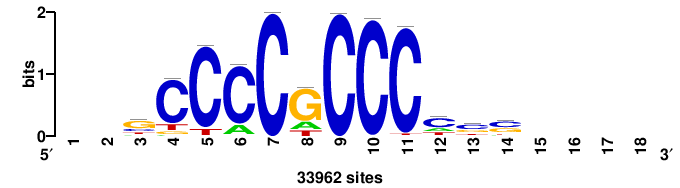
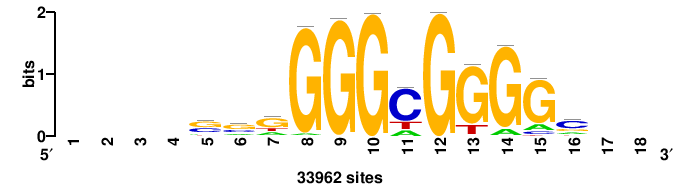
+yhmCGCCCMCGCAyt++ |
++ARTGCGKGGGCGKDR+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m347_MA0103_3 |
ZEB1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.65 |
20129 |
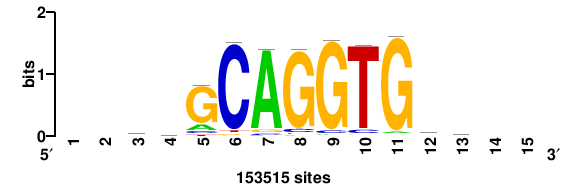
++SSSCAGGTGSS++ |
++ssCACCTGsss++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m26_MA0116_1 |
Znf423 |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
12.42 |
33 |
KMMMCCYWRGGKKSC |
gsmmCCyWrGGkkkm |
 |
 |
| JASPAR_2022_vertebrates_CORE_m669_MA1930_1 |
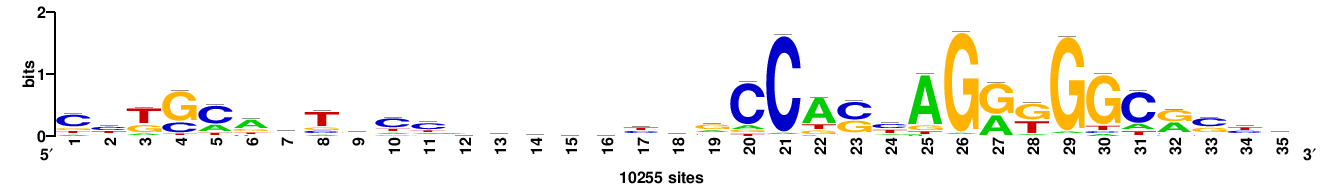
CTCF |
cluster_76 |
JASPAR_2022_vertebrates_CORE |
35 |
14.53 |
2576 |
YRSYGCCMYCTRSTGGCCRGRWGSKGGMACTGCAG |
cTGCAgTkccmscwycyggCCAsyAGrkGGCrsyr |
 |
 |
| JASPAR_2022_vertebrates_CORE_m804_MA0819_2 |
CLOCK |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.53 |
5744 |
+++araCACGTGb+++++ |
+++++VCACGTGTYT+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m825_MA1498_2 |
HOXA7 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.71 |
9611 |
++KTAATKRS+++ |
+++symATTAm++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m562_MA1634_1 |
BATF |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
7.61 |
47008 |
++++waTGACTCAth+++++ |
+++++DATGAGTCATW++++ |
 |
 |
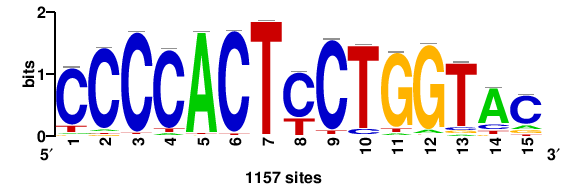
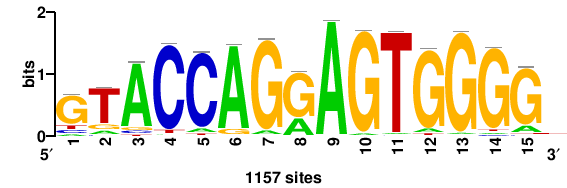
| JASPAR_2022_vertebrates_CORE_m550_MA1715_1 |
ZNF707 |
cluster_128 |
JASPAR_2022_vertebrates_CORE |
15 |
14.24 |
1157 |
CCCCACTYCTGGTAC |
GTACCAGRAGTGGGG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m161_MA0132_2 |
PDX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.87 |
7422 |
++YTAATKRS+++ |
+++symATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m58_MA0518_1 |
Stat4 |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
9.34 |
2873 |
++++yTTCyrRGAArysr |
YSRYTTCYYRGAAR++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m650_MA0745_2 |
SNAI2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.96 |
45816 |
+aygCACCTGTmry+ |
+RYKACAGGTGCRT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m205_MA0767_1 |
GCM2 |
cluster_52 |
JASPAR_2022_vertebrates_CORE |
11 |
7.63 |
2036 |
bATGCGGGyr+ |
+YRCCCGCATV |
 |
 |
| JASPAR_2022_vertebrates_CORE_m427_MA1523_1 |
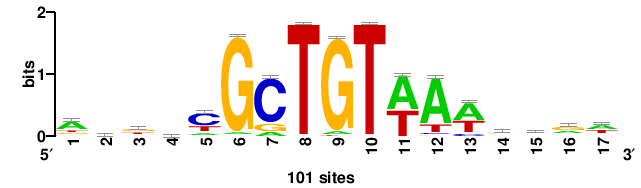
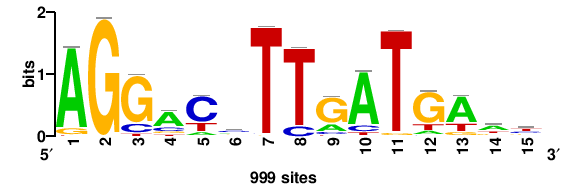
MSANTD3 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
6.17 |
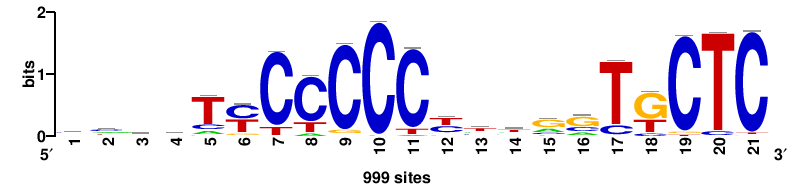
999 |
++KTGAGTGKAS++++ |
++++stmCACTCAm++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m218_MA0780_1 |
PAX3 |
cluster_35 |
JASPAR_2022_vertebrates_CORE |
10 |
10.69 |
10576 |
TAATYRATTA |
TAATyrATTA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m210_MA0498_2 |
MEIS1 |
cluster_34 |
JASPAR_2022_vertebrates_CORE |
17 |
5.50 |
94598 |
++++++CTGTCAW++++ |
++++wTGACAg++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m805_MA0821_2 |
HES5 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.20 |
14528 |
++++grCACGTGyc++++ |
++++GRCACGTGYC++++ |
 |
 |
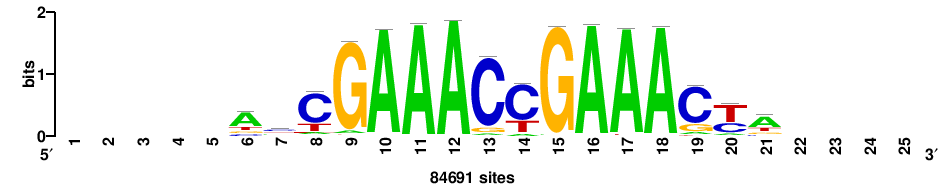
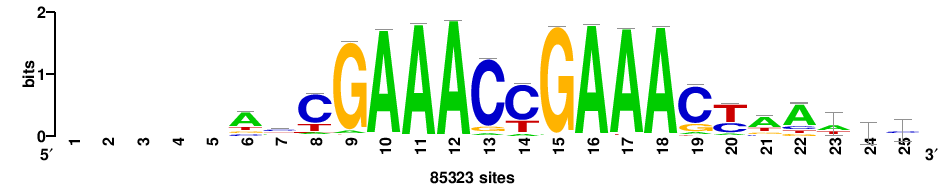
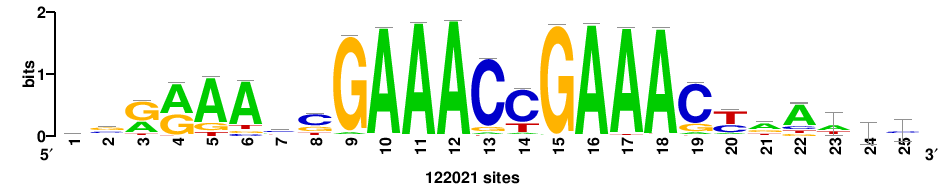
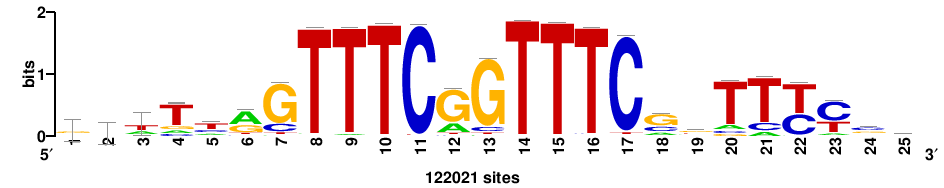
| JASPAR_2022_vertebrates_CORE_m208_MA0772_1 |
IRF7 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
12.22 |
3677 |
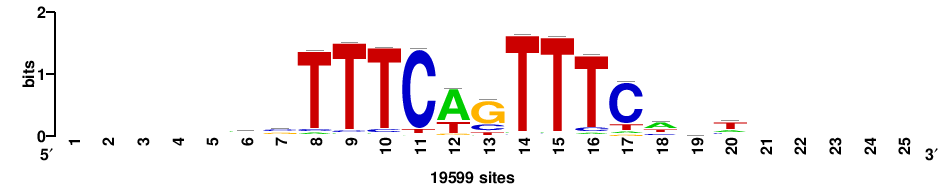
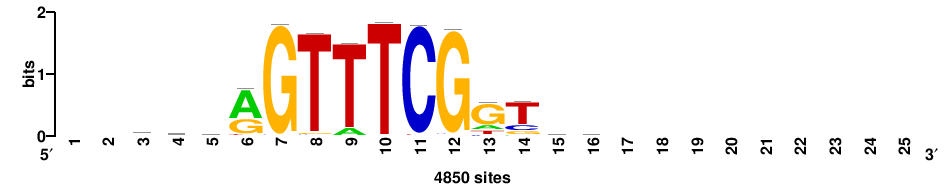
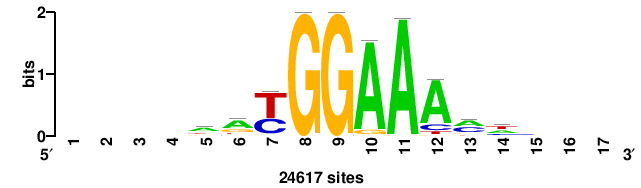
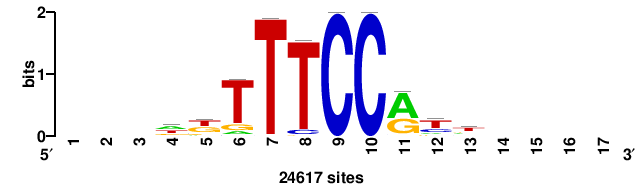
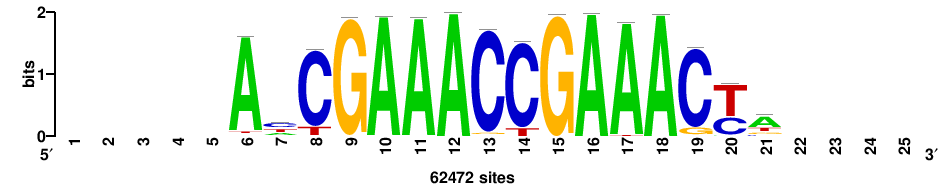
++++++ABTTTCRYTTTCGK+++++ |
+++++mCGAAAryGAAAvT++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m672_MA1933_1 |
ELK1::SREBF2 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
12.88 |
1315 |
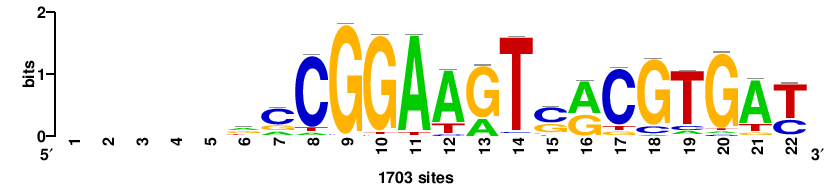
+++++rcCGGAAGTsrCGTGA+ |
+TCACGYSACTTCCGGY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m277_MA0850_1 |
FOXP3 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
5.96 |
283 |
+++++++TGTTTAY+++++ |
+++++rTAAACA+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m502_MA1627_1 |
Wt1 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.29 |
4691 |
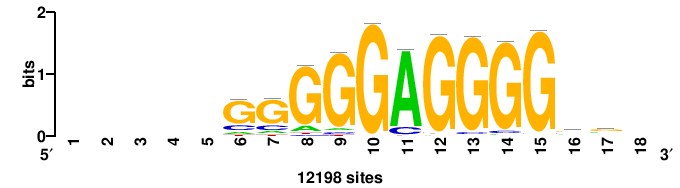
+GKGTGGGGGAGGRG+++ |
+++cycCTCCCCCACmc+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m654_MA0522_3 |
TCF3 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.10 |
31261 |
++svCACCTGCss++ |
++SSGCAGGTGBS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m134_MA0675_1 |
NKX6-2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.93 |
24187 |
++TTAATKRS+++ |
+++symATTAa++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m374_MA1148_1 |
PPARA::RXRA |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
8.61 |
1001 |
ATGACCTTTGACCYACWT+ |
+awgtrGGtcAaAGGTcAt |
 |
 |
| JASPAR_2022_vertebrates_CORE_m597_MA0836_2 |
CEBPD |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
8.33 |
21452 |
WWRTTGTGCAAYM++ |
++krTTGCACAAyww |
 |
 |
| JASPAR_2022_vertebrates_CORE_m279_MA0853_1 |
Alx4 |
cluster_22 |
JASPAR_2022_vertebrates_CORE |
18 |
9.35 |
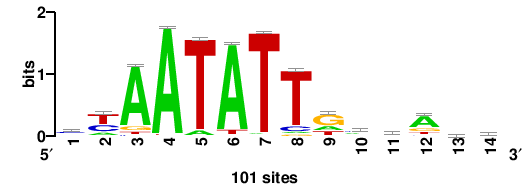
101 |
+cgcryTAATTArthasc |
GSTDAYTAATTARYGCG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m184_MA0472_2 |
EGR2 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
10.60 |
4311 |
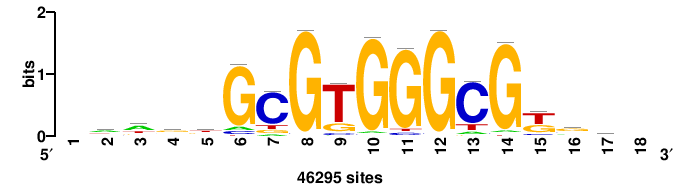
+++TGCGTGGGCGK++++ |
++++mCGCCCACGCa+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m500_MA1624_1 |
Stat5a |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
8.72 |
4514 |
+++DTTTCTTGGAAW+++ |
+++wTTCCAAGAAah+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m102_MA0475_2 |
FLI1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.20 |
54661 |
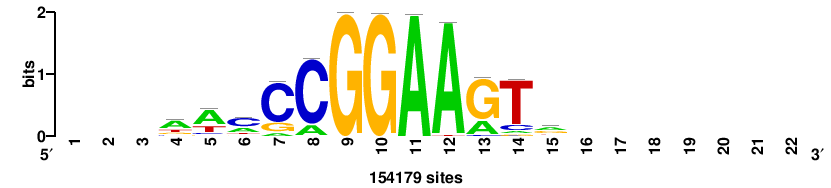
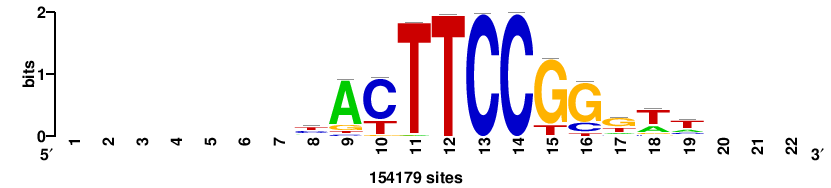
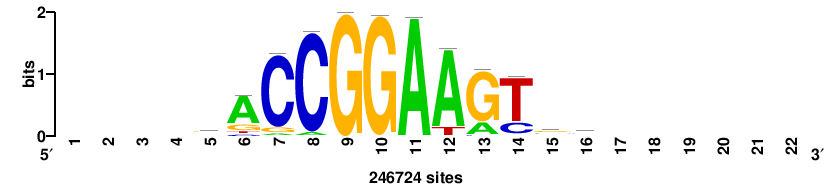
+++++ACCGGAArTg+++++++ |
+++++++CAYTTCCGGT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m158_MA0710_1 |
NOTO |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.64 |
33076 |
+vytMATTArs++ |
++SYTAATKARB+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m293_MA0866_1 |
SOX21 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
16.20 |
18296 |
++++AACAATkkyAkTGTT |
AACAMTRMMATTGTT++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m724_MA1985_1 |
ZNF669 |
cluster_14 |
JASPAR_2022_vertebrates_CORE |
16 |
11.00 |
1000 |
+AGYGGTCRTCRCCCC |
gGGgyGAyGACCrcT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m738_MA1999_1 |
Prdm5 |
cluster_94 |
JASPAR_2022_vertebrates_CORE |
15 |
8.12 |
740 |
ctGTTCTCCATCtss |
SSAGATGGAGAACAG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m786_MA0693_3 |
Vdr |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
6.72 |
38542 |
+++++++ywgAGTTCAtw++ |
++WATGAACTCWR+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m90_MA0630_1 |
SHOX |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.17 |
1000 |
++YYAATTAR+++ |
+++yTAATTrr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m113_MA0652_1 |
IRF8 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
15.60 |
7661 |
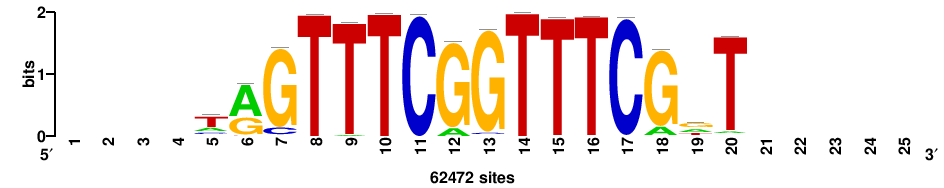
++++++yCGAAACCGAAACT+++++ |
+++++AGTTTCGGTTTCGR++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m678_MA1939_1 |
ERF::SREBF2 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
14.44 |
507 |
+++++ssCGGAArTCACGTGAy |
RTCACGTGAYTTCCGSS+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m579_MA1652_1 |
ZKSCAN5 |
cluster_74 |
JASPAR_2022_vertebrates_CORE |
14 |
9.38 |
3327 |
rgaGGArGTGAgrr |
YYCTCACYTCCTCY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m671_MA1932_1 |
ELK1::HOXB13 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
13.15 |
8465 |
+++++rCCGGAAGThrTAAAa+ |
+TTTTAYDACTTCCGGY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m10_MA0066_1 |
PPARG |
cluster_59 |
JASPAR_2022_vertebrates_CORE |
20 |
14.12 |
28 |
WGTRGGTCACSGTGACCYAS |
sTrGGTCAcsgTGACCyAcW |
 |
 |
| JASPAR_2022_vertebrates_CORE_m226_MA0790_1 |
POU4F1 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
9.10 |
6736 |
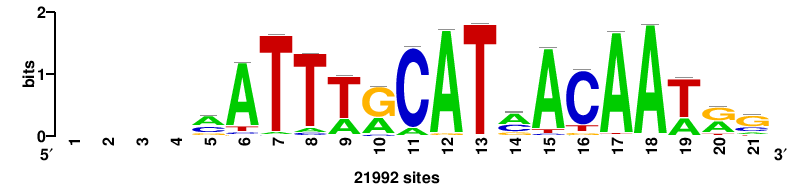
++++CATTAATWATKCAT+ |
+aTgmATwATTaATg++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m107_MA0648_1 |
GSC |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
6.68 |
8323 |
+syTAATCCsy+ |
+RSGGATTARS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m749_MA0087_2 |
Sox5 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
8.20 |
124 |
+rrsAACAATGGsm+++++ |
+++++KSCCATTGTTSYY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m508_MA0463_2 |
BCL6 |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
11.39 |
9142 |
wyGCTTTCkAGGAAyk++ |
++MRTTCCTMGAAAGCRW |
 |
 |
| JASPAR_2022_vertebrates_CORE_m479_MA1589_1 |
ZNF140 |
cluster_111 |
JASPAR_2022_vertebrates_CORE |
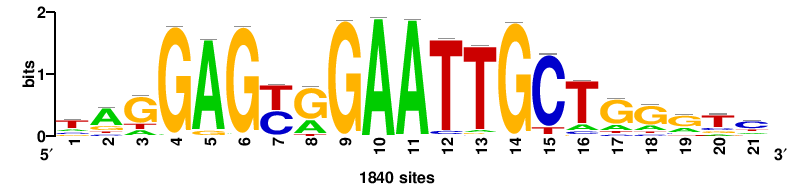
21 |
15.93 |
1840 |
tagGAGyrGAATTGCTgggtc |
GACCCAGCAATTCYRCTCCTA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m299_MA0875_1 |
BARX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
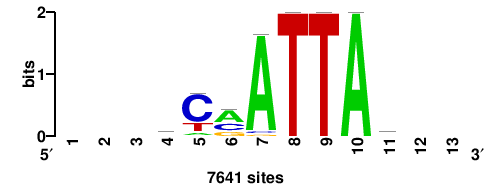
6.16 |
7641 |
++YTAATKGB+++ |
+++vcmATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m280_MA0854_1 |
Alx1 |
cluster_22 |
JASPAR_2022_vertebrates_CORE |
18 |
9.43 |
100 |
+cgmryTAATTArtymcy |
RGKRAYTAATTARYKCG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m761_MA0493_2 |
KLF1 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.24 |
999 |
+++CCMCRCCCM++++++ |
++++++kGGGyGkGG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m99_MA0027_2 |
EN1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.48 |
3593 |
++syAATTAg+++ |
+++CTAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m546_MA1711_1 |
ZNF343 |
cluster_126 |
JASPAR_2022_vertebrates_CORE |
17 |
13.39 |
59571 |
cCGCTTCmCcrCGGymv |
BKRCCGYGGKGAAGCGG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m358_MA1131_1 |
FOSL2::JUN |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
9.12 |
1001 |
++++grTGACGTmAt+++++ |
+++++ATKACGTCAYC++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m682_MA1943_1 |
ETV2::HOXB13 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
11.16 |
7799 |
+++++KTTWATRKWTCCGGT++ |
++aCCGGAwmyATwAAm+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m183_MA0731_1 |
BCL6B |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
20.66 |
384 |
+TGCTTTCTAGGAATTmm |
KKAATTCCTAGAAAGCA+ |
 |
 |
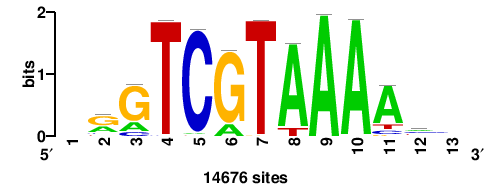
| JASPAR_2022_vertebrates_CORE_m523_MA0485_2 |
HOXC9 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
9.87 |
22481 |
++GTCGTAAAAy+ |
+RTTTTACGAC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m29_MA0130_1 |
ZNF354C |
cluster_131 |
JASPAR_2022_vertebrates_CORE |
6 |
6.21 |
16 |
mTCCAC |
GTGGAK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m475_MA1584_1 |
ZIC5 |
cluster_46 |
JASPAR_2022_vertebrates_CORE |
16 |
13.19 |
27639 |
mgACCCCCCGCtGyGm |
KCRCAGCGGGGGGTCK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m812_MA0877_3 |
BARHL1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
7.17 |
8627 |
++++++++TAAACGGT+++ |
+++acCGTTTA++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m397_MA1481_1 |
DRGX |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.43 |
18731 |
++yyAATTAs+++ |
+++STAATTRR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m123_MA0663_1 |
MLX |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
10.20 |
562 |
++++rTCACGTGAt++++ |
++++ATCACGTGAY++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m64_MA0258_2 |
ESR2 |
cluster_59 |
JASPAR_2022_vertebrates_CORE |
20 |
9.09 |
8243 |
++RGGTCASKSTGACCY+++ |
+++rGGTCAsmstGaCCy++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m708_MA1969_1 |
THRA |
cluster_58 |
JASPAR_2022_vertebrates_CORE |
24 |
14.38 |
182 |
RGWRAGGTCAYRTGAGGWCACAGG |
cctgTGwCCTCAyrTGACCTywcy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m287_MA0525_2 |
TP63 |
cluster_50 |
JASPAR_2022_vertebrates_CORE |
18 |
16.87 |
4021 |
GACATGTYMCRACATGTY |
rACATGTygkrACATGTc |
 |
 |
| JASPAR_2022_vertebrates_CORE_m45_MA0468_1 |
DUX4 |
cluster_38 |
JASPAR_2022_vertebrates_CORE |
16 |
10.41 |
38217 |
++TGATTRRRTTA+++ |
+++TAAyyyAATCA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m87_MA0626_1 |
Npas2 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.98 |
1000 |
++++dsCACGTGtm++++ |
++++KACACGTGSH++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m424_MA1520_1 |
MAF |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.74 |
3922 |
++wTGCTGAsTCAGCAw+++ |
+++WTGCTGASTCAGCAW++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m453_MA1557_1 |
SMAD5 |
cluster_15 |
JASPAR_2022_vertebrates_CORE |
12 |
9.39 |
7598 |
yGTCTAGACa++ |
++TGTCTAGACR |
 |
 |
| JASPAR_2022_vertebrates_CORE_m467_MA1575_1 |
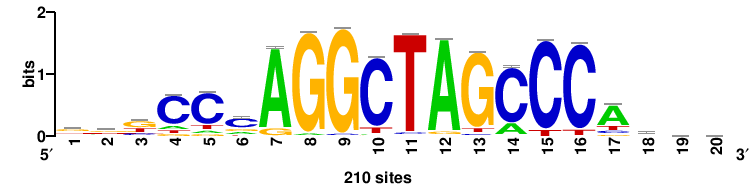
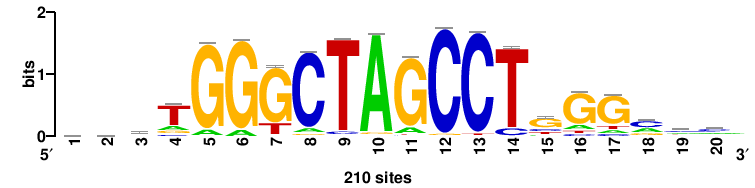
THRB |
cluster_136 |
JASPAR_2022_vertebrates_CORE |
19 |
12.37 |
6681 |
gTGwCCTyrhyrAGGwCAc |
GTGWCCTYRDYRAGGWCAC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m773_MA0624_2 |
Nfatc1 |
cluster_44 |
JASPAR_2022_vertebrates_CORE |
17 |
7.93 |
6191 |
+++saaTGGAAAaww++ |
++WWTTTTCCATTS+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m582_MA1655_1 |
ZNF341 |
cluster_57 |
JASPAR_2022_vertebrates_CORE |
15 |
7.50 |
18728 |
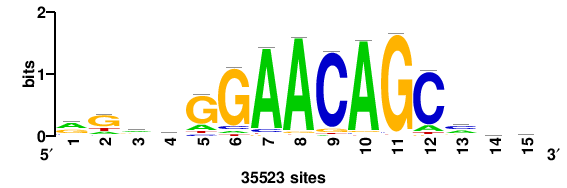
STGGCTGTTCYC+++ |
+++grGAACAGCcas |
 |
 |
| JASPAR_2022_vertebrates_CORE_m48_MA0479_1 |
FOXH1 |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
9.54 |
8211 |
+TGTGKATTSVR |
ybsAATMCACA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m647_MA0684_2 |
RUNX3 |
cluster_60 |
JASPAR_2022_vertebrates_CORE |
14 |
7.15 |
27377 |
+DWTTGAGGTTWT+ |
+awAACCTCAawh+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m662_MA0093_3 |
USF1 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
9.08 |
55334 |
++DRGTCACATGACYH++ |
++drGTCATGTGACyh++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m340_MA1118_1 |
SIX1 |
cluster_33 |
JASPAR_2022_vertebrates_CORE |
16 |
8.31 |
1108 |
+KMTCAGGTTWC++++ |
++++GwAACCTGAkm+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m721_MA1982_1 |
ZNF574 |
cluster_77 |
JASPAR_2022_vertebrates_CORE |
16 |
10.78 |
998 |
+SGGCCKCTCTAGSMS |
sksCTAGAGmGGCcs+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m531_MA0623_2 |
NEUROG1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
9.55 |
3772 |
++raCATATGky+++ |
+++RMCATATGTY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m259_MA0058_3 |
MAX |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.93 |
7263 |
++++RGCACGTGKT++++ |
++++amCACGTGcy++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m820_MA1472_2 |
Bhlha15 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.26 |
21916 |
+SMACAGCTGTKS++ |
++smACAGCTGTks+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m409_MA1502_1 |
HOXB8 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.73 |
7937 |
++KTAATKRC+++ |
+++gymATTAm++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m55_MA0504_1 |
NR2C2 |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
11.26 |
395 |
++mgrGGTCArAGGTCA++ |
++TGACCTYTGACCYCK++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m764_MA0507_2 |
POU2F2 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
12.54 |
11662 |
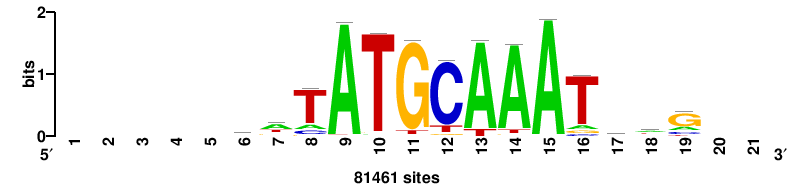
++CTMATTTGCATAWT+++++ |
+++++awTATGCAAATkAg++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m413_MA1506_1 |
HOXD10 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
10.17 |
6159 |
+rGTCGTAAAAh+ |
+DTTTTACGACY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m118_MA0657_1 |
KLF13 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
14.22 |
6580 |
rtRmCACGCCCCYTTttg |
CAAAARGGGGCGTGKYAY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m152_MA0698_1 |
ZBTB18 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
10.69 |
21996 |
+VMACATCTGGATD+ |
+hatCCAGATGTkb+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m330_MA0104_4 |
MYCN |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
8.21 |
6984 |
+++SGCCACGTGGCS+++ |
+++sgCCACGTGGcs+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m513_MA0766_2 |
GATA5 |
cluster_11 |
JASPAR_2022_vertebrates_CORE |
15 |
7.93 |
19252 |
+cwGATAAsrm++++ |
++++KYSTTATCWG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m537_MA0746_2 |
SP3 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
10.30 |
7826 |
+RGKGGGCGTGGYS++++ |
++++srCCACGCCCmCy+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m544_MA1709_1 |
ZIM3 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
10.92 |
17220 |
+++++++TRGGTTTCTGTTKYT+++ |
+++armAACAGAAAcCya+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m375_MA1149_1 |
RARA::RXRG |
cluster_31 |
JASPAR_2022_vertebrates_CORE |
18 |
9.23 |
1001 |
TGACCYYTMCWTGACCYY |
rrGGTCAwgkarrGGtCA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m685_MA1946_1 |
ETV5::FOXI1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.85 |
661 |
+GtMAACMGGAwGy++++++++ |
++++++++RCWTCCKGTTKAC+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m690_MA1951_1 |
FOS |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
12.06 |
2303 |
+++YGATGACGTCATCV+++ |
+++bgATGACGTCATCr+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m700_MA1961_1 |
PATZ1 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
10.21 |
999 |
++SCCCCKCCCCSS++++ |
++++ssGGGGmGGGGs++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m698_MA1959_1 |
KLF7 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
8.93 |
998 |
+++CCMCGCCCH++++++ |
++++++dGGGCGKgG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m119_MA0658_1 |
LHX6 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.60 |
5757 |
+rcTAATTAry++ |
++RYTAATTAGY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m660_MA0814_2 |
TFAP2C |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
8.03 |
60819 |
+SBCGCCTCAGGCCV |
bggCCTGAGGCgvs+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m741_MA2003_1 |
NKX2-4 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
6.40 |
3369 |
++++cCACTTsAvm++ |
++KBTSAAGTGG++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m419_MA1514_1 |
KLF17 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
11.54 |
39844 |
+RAKGGGTGCGTGGKK++ |
++mmCCACgCaCCCmty+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m46_MA0476_1 |
FOS |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
9.45 |
29396 |
++++dvTGAsTCATb+++++ |
+++++VATGASTCABH++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m731_MA1992_1 |
Ikzf3 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.37 |
1183 |
+++rsrCAGGAAGTGggg++++ |
++++CCCCACTTCCTGYSY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m556_MA1721_1 |
ZNF93 |
cluster_75 |
JASPAR_2022_vertebrates_CORE |
18 |
11.23 |
2388 |
+gGyrkCrGCAGCGGyg+ |
+CRCCGCTGCYGMYRCC+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m411_MA1504_1 |
HOXC4 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.54 |
12058 |
++YTAATKRY+++ |
+++rYmATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m2_MA0006_1 |
Ahr::Arnt |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
6.61 |
24 |
++++++yGCGTG++++++ |
++++++CACGCR++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m32_MA0142_1 |
Pou5f1::Sox2 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
10.26 |
1369 |
+++++ATTTGCATRWSAAWR+ |
+ywTTswyATGCAaAt+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m747_MA0079_5 |
SP1 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.93 |
999 |
+++CYCCKCCCC++++++ |
++++++GGGGMGGrG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m474_MA1583_1 |
ZFP57 |
cluster_102 |
JASPAR_2022_vertebrates_CORE |
13 |
8.09 |
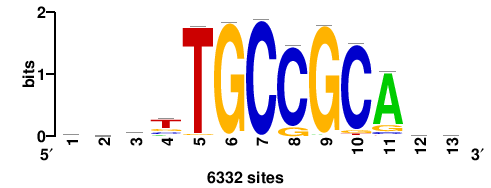
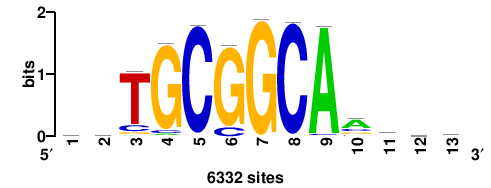
6332 |
bsrtTGCCGCAsb |
VSTGCGGCAAYSV |
 |
 |
| JASPAR_2022_vertebrates_CORE_m828_MA1524_2 |
Msgn1 |
cluster_84 |
JASPAR_2022_vertebrates_CORE |
21 |
10.10 |
542 |
RYCRCCATTTGTYYYY+++++ |
+++++rrrraCAAATGGygry |
 |
 |
| JASPAR_2022_vertebrates_CORE_m806_MA0828_2 |
SREBF2 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
12.28 |
14825 |
++WTRTCACGTGAYAW++ |
++wtrTCACGTGAyaw++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m103_MA0042_2 |
FOXI1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.05 |
6190 |
+++++++TGTTTAC+++++ |
+++++GTAAACA+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m315_MA0897_1 |
Hmx2 |
cluster_22 |
JASPAR_2022_vertebrates_CORE |
18 |
9.97 |
101 |
+acaAgCAATTAAwgvrT |
AYBCWTTAATTGCTTGT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m759_MA0480_2 |
Foxo1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
6.52 |
14116 |
+++++rwGTAAACAva+++ |
+++TBTGTTTACWY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m670_MA1931_1 |
ELK1::HOXA1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
12.05 |
123 |
+++++aCCGGAAGTAAtTa+++ |
+++TAATTACTTCCGGT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m534_MA0075_3 |
PRRX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.64 |
11044 |
++cyAATTAr+++ |
+++YTAATTRG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m117_MA0656_1 |
JDP2 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
12.14 |
27142 |
++++GRTGACGTCATC++++ |
++++gATGACGTCAyc++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m169_MA0720_1 |
Shox2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.41 |
20191 |
++yyAATTAr+++ |
+++YTAATTRR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m653_MA0829_2 |
SREBF1 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
9.37 |
7141 |
++rrATCACsTGATyy++ |
++RRATCASGTGATYY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m710_MA1971_1 |
ZBED2 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
7.55 |
1297 |
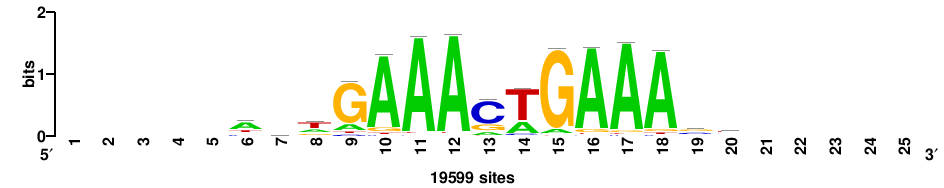
+++++++++svmmCGAAACCvvv++ |
++BBBGGTTTCGKKBS+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m253_MA0461_2 |
Atoh1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.89 |
1281 |
++RMCATATGKY+++ |
+++rmCATATGky++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m452_MA1556_1 |
RXRG |
cluster_20 |
JASPAR_2022_vertebrates_CORE |
14 |
12.42 |
2528 |
RRGGTCRTGACCYY |
rrGGTCAyGACCyy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m630_MA0620_3 |
MITF |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
9.04 |
63527 |
wwwrrTCACGTGAyywww |
WWWRRTCACGTGAYYWWW |
 |
 |
| JASPAR_2022_vertebrates_CORE_m567_MA1639_1 |
MEIS1 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
8.25 |
6736 |
++rhcATmAATCAww++++ |
++++WWTGATTKATGDY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m92_MA0634_1 |
ALX3 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.17 |
9897 |
+yyyAATTArm++ |
++KYTAATTRRR+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m781_MA0666_2 |
MSX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.11 |
7706 |
++syAATTAr+++ |
+++YTAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m676_MA1937_1 |
ERF::HOXB13 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.40 |
3962 |
+++++rccGGAaryhrTwAaa+ |
+TTTWAYDRYTTCCGGY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m827_MA1518_2 |
Lhx1 |
cluster_22 |
JASPAR_2022_vertebrates_CORE |
18 |
7.84 |
454 |
RRGTGCTAATTAGCACYY |
rrgtgcTAATTAgcacyy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m18_MA0084_1 |
SRY |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
6.37 |
28 |
+++++++WTTGTTWWM+++ |
+++kwwaACAAw+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m593_MA0833_2 |
ATF4 |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
10.37 |
33276 |
rraTGATGCAATmw+ |
+WKATTGCATCATYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m529_MA0702_2 |
LMX1A |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
8.61 |
7958 |
++yTAATTAm+++ |
+++KTAATTAR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m389_MA1470_1 |
BACH2 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
15.70 |
436 |
awAbsATGACGTSATsmTWt |
AWAKSATSACGTCATSVTWT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m359_MA1132_1 |
JUN::JUNB |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
6.74 |
1001 |
++++kaTGACkCAt++++++ |
++++++ATGMGTCATM++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m480_MA1592_1 |
ZNF274 |
cluster_114 |
JASPAR_2022_vertebrates_CORE |
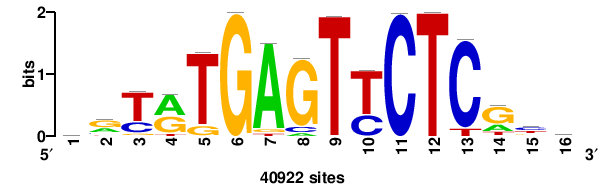
16 |
11.79 |
40922 |
rryrTGAGTyCTCgyk |
MRCGAGRACTCAYRYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m220_MA0783_1 |
PKNOX2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
14.71 |
1423 |
+TGACASCTGTCA++ |
++TGACAGsTGTCA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m765_MA0509_3 |
RFX1 |
cluster_25 |
JASPAR_2022_vertebrates_CORE |
17 |
16.01 |
19362 |
+SGTTRCCATGGCAACS |
sGTTGCCATGGyAACs+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m532_MA0842_2 |
NRL |
cluster_51 |
JASPAR_2022_vertebrates_CORE |
16 |
10.48 |
31751 |
++aawwrTGCTGACg+ |
+CGTCAGCAYWWTT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m616_MA1104_2 |
GATA6 |
cluster_11 |
JASPAR_2022_vertebrates_CORE |
15 |
7.72 |
79444 |
WAAGATAAGAWWW++ |
++wwwTCTTATCTtw |
 |
 |
| JASPAR_2022_vertebrates_CORE_m44_MA0146_2 |
Zfx |
cluster_79 |
JASPAR_2022_vertebrates_CORE |
14 |
9.10 |
481 |
sssgsCbvGGCCTs |
SAGGCCBVGSCSSS |
 |
 |
| JASPAR_2022_vertebrates_CORE_m449_MA1553_1 |
RARG |
cluster_20 |
JASPAR_2022_vertebrates_CORE |
14 |
11.06 |
4618 |
rAGGTCATGACCTy |
RAGGTCATGACCTY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m819_MA1467_2 |
Atoh1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.84 |
21477 |
+KRCCATCTGBY+++ |
+++rvCAGATGGym+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m585_MA1683_1 |
FOXA3 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
6.98 |
48937 |
+++++rwGTAAACAww+++ |
+++WWTGTTTACWY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m165_MA0716_1 |
PRRX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.70 |
28467 |
++yyAATTAr+++ |
+++YTAATTRR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m235_MA0510_2 |
RFX5 |
cluster_25 |
JASPAR_2022_vertebrates_CORE |
17 |
14.18 |
3094 |
+sGTTrCYATGGyAACs |
SGTTRCCATRGYAACS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m247_MA0811_1 |
TFAP2B |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
7.53 |
4959 |
++tgCCyyvrGGCa+ |
+TGCCYBRRGGCA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m105_MA0646_1 |
GCM1 |
cluster_52 |
JASPAR_2022_vertebrates_CORE |
11 |
9.03 |
34329 |
cATGCGGGTac |
GTACCCGCATG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m472_MA1580_1 |
ZBTB32 |
cluster_124 |
JASPAR_2022_vertebrates_CORE |
10 |
10.19 |
3572 |
tGTACAGTaw |
WTACTGTACA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m192_MA0741_1 |
KLF16 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.12 |
3053 |
++gmCMCgCCCMC+++++ |
+++++GKGGGCGKGKC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m594_MA0835_2 |
BATF3 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.09 |
11965 |
++++waTGACTCAtw+++++ |
+++++WATGAGTCATW++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m104_MA0033_2 |
FOXL1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
7.37 |
5465 |
+++++++TGTTTAY+++++ |
+++++rTAAACA+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m776_MA0642_2 |
EN2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.31 |
3367 |
+++TAATTRSK++ |
++msyAATTA+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m448_MA1552_1 |
RARB |
cluster_20 |
JASPAR_2022_vertebrates_CORE |
14 |
11.15 |
609 |
rAGGTCrTGACCTy |
RAGGTCAYGACCTY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m692_MA1953_1 |
FOXO1::ELF1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.32 |
440 |
+rtmAACAGGAAgtd+++++++ |
+++++++HACTTCCTGTTKAY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m6_MA0031_1 |
FOXD1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.27 |
20 |
+++++++WTGTTTAC++++ |
++++GTAAACAw+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m587_MA1726_1 |
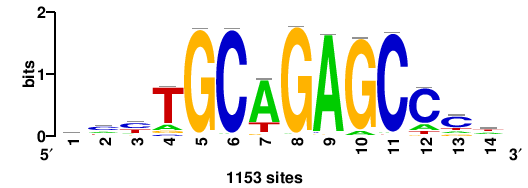
ZNF331 |
cluster_98 |
JASPAR_2022_vertebrates_CORE |
14 |
9.43 |
1153 |
mcyTGCAGAGCCcw |
WGGGCTCTGCARGK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m382_MA1420_1 |
IRF5 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
10.64 |
508 |
++++++cCGAAACCGAAmCy+++++ |
+++++RGKTTCGGTTTCGG++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m275_MA0848_1 |
FOXO4 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
7.39 |
7060 |
+++++++GTAAACA+++++ |
+++++TGTTTAC+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m349_MA1125_1 |
ZNF384 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
9.08 |
30433 |
+TTTTTTTTKWWW++ |
++wwwmAAAAAAaa+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m296_MA0028_2 |
ELK1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.92 |
67234 |
+++++ACCGGAAGTr+++++++ |
+++++++YACTTCCGGT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m604_MA0473_3 |
ELF1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.78 |
47770 |
+++aamCAGGAAGTGrv+++++ |
+++++BYCACTTCCTGKTT+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m610_MA0047_3 |
FOXA2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
7.43 |
262153 |
+++++awGTAAACAww+++ |
+++WWTGTTTACWT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m74_MA0601_1 |
Arid3b |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.14 |
1001 |
++TTWATTAATWT |
awaTTAATwaa++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m202_MA0098_3 |
ETS1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.42 |
3774 |
+++++ACCGGAWrTr+++++++ |
+++++++YAYWTCCGGT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m470_MA1578_1 |
VEZF1 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
8.06 |
999 |
+++WARKGGGGGG+++++ |
+++++CCCCCCmytw+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m792_MA0740_2 |
KLF14 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.45 |
998 |
+++CCMCGCCCM++++++ |
++++++kGGGCGKGG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m560_MA1631_1 |
ASCL1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.20 |
34356 |
+cdgCACCTGCysc+ |
+GSRGCAGGTGCHG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m600_MA0471_2 |
E2F6 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
8.04 |
33098 |
++grrGGCGGGAArs+++ |
+++SYTTCCCGCCYYC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m778_MA0650_3 |
Hoxa13 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
7.74 |
17248 |
+wgCCAATAAAaw++ |
++WTTTTATTGGCW+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m250_MA0524_2 |
TFAP2C |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
7.25 |
8544 |
++tGCCyyrrGGCa+ |
+TGCCYYRRGGCA++ |
 |
 |
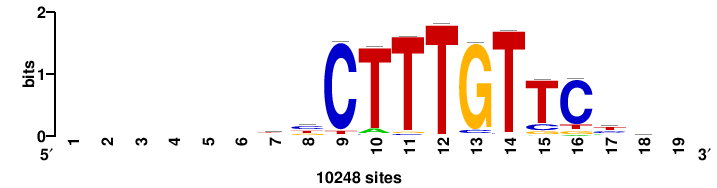
| JASPAR_2022_vertebrates_CORE_m811_MA0869_2 |
Sox11 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
8.26 |
10248 |
+BYCTTTGTTCYY++++++ |
++++++rrGAACAAAGrv+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m445_MA1548_1 |
PLAGL2 |
cluster_62 |
JASPAR_2022_vertebrates_CORE |
14 |
8.40 |
999 |
+++AGGGGGCCCA+ |
+tGGGCCCCCt+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m229_MA0793_1 |
POU6F2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
8.11 |
4277 |
asCTmATTAw+++ |
+++WTAATKAGST |
 |
 |
| JASPAR_2022_vertebrates_CORE_m436_MA1536_1 |
NR2C2 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
5.21 |
25267 |
+++++++++rrGGTCRh+++ |
+++DYGACCYY+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m83_MA0618_1 |
LBX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.04 |
1000 |
++CYARTTAA+++ |
+++tTAAYTRG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m360_MA1133_1 |
JUN::JUNB |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
7.63 |
1001 |
++++krTGACGTCATb++++ |
++++VATGACGTCAYM++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m622_MA0490_2 |
JUNB |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.74 |
79856 |
+++wdATGACTCATwh++++ |
++++DWATGAGTCATHW+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m519_MA0158_2 |
HOXA5 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.05 |
4843 |
++STAATKRS+++ |
+++symATTAs++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m82_MA0615_1 |
Gmeb1 |
cluster_106 |
JASPAR_2022_vertebrates_CORE |
17 |
5.80 |
101 |
smkkkkRCGYmmgayrs |
SYRTCKKRCGYMMMMKS |
 |
 |
| JASPAR_2022_vertebrates_CORE_m715_MA1976_1 |
ZNF320 |
cluster_78 |
JASPAR_2022_vertebrates_CORE |
27 |
12.97 |
999 |
rGkGGGrcCmrGGGGMmwrtGmgrsks |
SMSYCKCAYWKKCCCCYKGGYCCCMCY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m575_MA1648_1 |
TCF12 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
6.68 |
61876 |
++ssCACCTGCys++ |
++SRGCAGGTGSS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m68_MA0140_2 |
GATA1::TAL1 |
cluster_120 |
JASPAR_2022_vertebrates_CORE |
18 |
9.91 |
4955 |
cTTATCwskgagrarCAG |
CTGYTYCTCMSWGATAAG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m228_MA0792_1 |
POU5F1B |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
7.32 |
15038 |
+++++ATTWRCATA+++++++ |
+++++++TATGywAAT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m94_MA0635_1 |
BARHL2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
5.89 |
21181 |
++++++vhTAAAyGrt+++ |
+++AYCRTTTADB++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m483_MA1596_1 |
ZNF460 |
cluster_65 |
JASPAR_2022_vertebrates_CORE |
16 |
14.12 |
1612 |
rCCTCmGCCTCCCrrG |
CYYGGGAGGCKGAGGY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m264_MA0834_1 |
ATF7 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
12.00 |
27156 |
+++crATGACGTCAYmv+++ |
+++BKRTGACGTCATYG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m115_MA0654_1 |
ISX |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.34 |
2142 |
++yyAATTAr+++ |
+++YTAATTRR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m25_MA0115_1 |
NR1H2::RXRA |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
19.32 |
25 |
++GTTGACCTTTGACCTTT |
AAAGGTCAAAGGTCAAc++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m15_MA0073_1 |
RREB1 |
cluster_135 |
JASPAR_2022_vertebrates_CORE |
20 |
15.44 |
11 |
MCMCmAMmCAmCMmCmmmsm |
KSKKKGKKGKTGKKTKGKGK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m643_MA0681_2 |
PHOX2B |
cluster_38 |
JASPAR_2022_vertebrates_CORE |
16 |
10.39 |
53236 |
WWWTAATTTGATTAWW |
wwTAATCAAATTAwww |
 |
 |
| JASPAR_2022_vertebrates_CORE_m511_MA0470_2 |
E2F4 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
10.17 |
20375 |
+wTtTGGCGCCawww+++ |
+++WWWTGGCGCCAAAW+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m516_MA1099_2 |
HES1 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.60 |
34463 |
++++ggCrCGTGbc++++ |
++++GVCACGYGCC++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m303_MA0882_1 |
DLX6 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.77 |
3223 |
++syAATTAm+++ |
+++KTAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m493_MA1615_1 |
Plagl1 |
cluster_62 |
JASPAR_2022_vertebrates_CORE |
14 |
7.88 |
14153 |
+ssctGGGGCCAss |
SSTGGCCCCAGSS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m410_MA1503_1 |
HOXB9 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
9.29 |
2824 |
++gTCRTAAAat+ |
+ATTTTAYGAC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m673_MA1934_1 |
ERF::FIGLA |
cluster_40 |
JASPAR_2022_vertebrates_CORE |
16 |
12.69 |
18732 |
RMCASSTGYWTCCGGY |
rcCGGAwrCAssTGky |
 |
 |
| JASPAR_2022_vertebrates_CORE_m538_MA0632_2 |
TCFL5 |
cluster_67 |
JASPAR_2022_vertebrates_CORE |
15 |
8.21 |
161954 |
+++kCrCGCGCmc++ |
++GKGCGCGYGM+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m343_MA1120_1 |
SOX13 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
7.04 |
4552 |
+++raACAATGgmw+++++ |
+++++WKCCATTGTTY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m260_MA0825_1 |
MNT |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.74 |
713 |
++++RGCACGTGBT++++ |
++++avCACGTGcy++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m655_MA0830_2 |
TCF4 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.50 |
29465 |
+cggCACCTGccss+ |
+SSGGCAGGTGCCG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m497_MA1620_1 |
Ptf1A |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.60 |
10291 |
+rmaCACCTGtky++ |
++RMACAGGTGTKY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m569_MA1641_1 |
MYF5 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.19 |
11210 |
+gvaCAGCTGtbc++ |
++GVACAGCTGTBC+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m172_MA0724_1 |
VENTX |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.66 |
36776 |
++YTAATYGKT++ |
++AmCrATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m455_MA1559_1 |
SNAI3 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
9.30 |
11542 |
++TRCACCTGYT+++ |
+++arCAGGTGya++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m782_MA0668_2 |
Neurod2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.53 |
32957 |
BSRCCATCTGTYCYY |
rrgraCAGATGGysv |
 |
 |
| JASPAR_2022_vertebrates_CORE_m380_MA1418_1 |
IRF3 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
16.61 |
42165 |
YRGTTTCGGTTTCCKTTYYSS++++ |
++++ssrrAAmGGAAACCGAAACyr |
 |
 |
| JASPAR_2022_vertebrates_CORE_m206_MA0770_1 |
HSF2 |
cluster_16 |
JASPAR_2022_vertebrates_CORE |
15 |
11.93 |
4544 |
++TTCTAGAAyrTTC |
GAAYRTTCTAGAA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m386_MA1466_1 |
ATF6 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
13.23 |
1817 |
++tgrTGACGTGGCAs++ |
++STGCCACGTCAYCA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m716_MA1977_1 |
ZNF324 |
cluster_71 |
JASPAR_2022_vertebrates_CORE |
23 |
11.73 |
999 |
dmswrgmAGmyAAGGATGGYT++ |
++ARCCATCCTTRKCTKCYWSKH |
 |
 |
| JASPAR_2022_vertebrates_CORE_m316_MA0898_1 |
Hmx3 |
cluster_22 |
JASPAR_2022_vertebrates_CORE |
18 |
9.67 |
101 |
+asaAgCAATTAAwrrat |
ATYYWTTAATTGCTTST+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m371_MA1145_1 |
FOSL2::JUND |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
9.15 |
1001 |
++GCGRTGACGTMAYMM+++ |
+++kkrTkACGTCAycgc++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m234_MA0797_1 |
TGIF2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
12.74 |
15077 |
+TGACAGsTGTCA++ |
++TGACASCTGTCA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m646_MA0508_3 |
PRDM1 |
cluster_113 |
JASPAR_2022_vertebrates_CORE |
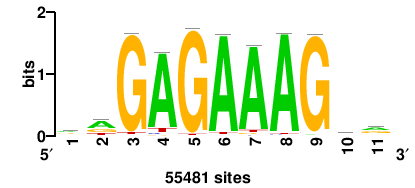
11 |
8.13 |
55481 |
ywCTTTCTCty |
RAGAGAAAGWR |
 |
 |
| JASPAR_2022_vertebrates_CORE_m168_MA0719_1 |
RHOXF1 |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
5.67 |
14231 |
++KGGMTYAW++ |
++wTrAkCCm++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m649_MA0744_2 |
SCRT2 |
cluster_37 |
JASPAR_2022_vertebrates_CORE |
16 |
9.02 |
65610 |
rrwkCAACAGGTggtw |
WACCACCTGTTGMWYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m725_MA1986_1 |
ZNF692 |
cluster_62 |
JASPAR_2022_vertebrates_CORE |
14 |
8.48 |
10745 |
+++SSTGGGCCCM+ |
+kgGGCCCAss+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m517_MA0616_2 |
HES2 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
8.50 |
12672 |
++++GgCACGTGyc++++ |
++++GRCACGTGCC++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m750_MA0095_3 |
Yy1 |
cluster_84 |
JASPAR_2022_vertebrates_CORE |
21 |
8.13 |
43348 |
++wmCAAAATGGck+++++++ |
+++++++MGCCATTTTGKW++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m24_MA0111_1 |
Spz1 |
cluster_85 |
JASPAR_2022_vertebrates_CORE |
11 |
8.25 |
12 |
RGGGTwwCAGc |
GCTGWWACCCY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m38_MA0155_1 |
INSM1 |
cluster_62 |
JASPAR_2022_vertebrates_CORE |
14 |
10.30 |
24 |
TGymwGGGGkcr++ |
++YGMCCCCWKRCA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m175_MA0112_3 |
ESR1 |
cluster_59 |
JASPAR_2022_vertebrates_CORE |
20 |
15.62 |
755 |
+MAGGTCAYSGTGACCYT++ |
++arGGTCAcsrTGACCTk+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m766_MA0514_2 |
Sox3 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
7.02 |
11978 |
+++raACAATGgma+++++ |
+++++TKCCATTGTTY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m471_MA1579_1 |
ZBTB26 |
cluster_16 |
JASPAR_2022_vertebrates_CORE |
15 |
8.02 |
1627 |
RKYTTTTCTGGAGHA |
tdcTCCAGAAaarmy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m141_MA0686_1 |
SPDEF |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.03 |
6861 |
++++YACATCCGGKT+++++++ |
+++++++AmCCGGATgTr++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m730_MA1991_1 |
Hnf1A |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
7.96 |
1249 |
++MVASATCAAAGGVV+++ |
+++bbcCTTTGATstbk++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m306_MA0887_1 |
EVX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.36 |
11702 |
+gvtaATTAsc++ |
++GSTAATTABC+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m739_MA2001_1 |
Six4 |
cluster_33 |
JASPAR_2022_vertebrates_CORE |
16 |
6.88 |
4291 |
+KVTCAGGTTWC++++ |
++++gwAACCTGAbm+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m73_MA0599_1 |
KLF5 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
8.91 |
13611 |
++DGGGYGKGGC++++++ |
++++++gCCMCrCCCh++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m438_MA1538_1 |
NR2F1 |
cluster_14 |
JASPAR_2022_vertebrates_CORE |
16 |
12.75 |
4036 |
rrGGTCRtTGACCYy+ |
+RRGGTCAAYGACCYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m425_MA1521_1 |
MAFA |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.49 |
10826 |
++dTGCTGAsTCAGCah+++ |
+++DTGCTGASTCAGCAH++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m801_MA0798_3 |
RFX3 |
cluster_25 |
JASPAR_2022_vertebrates_CORE |
17 |
16.07 |
2560 |
+sGTTrCCATGGCAACs |
SGTTGCCATGGYAACS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m626_MA0700_2 |
LHX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.35 |
6333 |
WWTAATTGGWW++ |
++wwcCAATTAww |
 |
 |
| JASPAR_2022_vertebrates_CORE_m324_MA0914_1 |
ISL2 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
5.25 |
11628 |
++++YTAAKTGC++++ |
++++gCAmTTAr++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m378_MA1152_1 |
SOX15 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
6.58 |
1001 |
++ARAACAAWRR+++++++ |
+++++++yywTTGTtyt++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m181_MA0729_1 |
RARA |
cluster_47 |
JASPAR_2022_vertebrates_CORE |
19 |
19.50 |
855 |
+MMTTGACCTTTTGACCTY |
rAGGTCAAAAGGTCAAkk+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m57_MA0517_1 |
STAT1::STAT2 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
11.36 |
620 |
+++++++RGRAAMYGAAACTRA+++ |
+++tyAGTTTCrkTTYCy+++++++ |
 |
 |
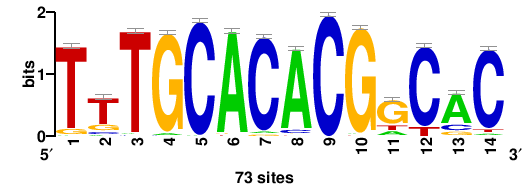
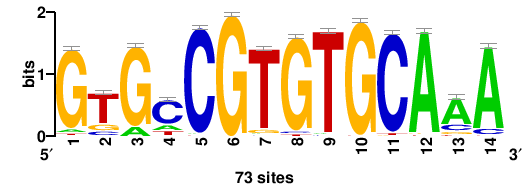
| JASPAR_2022_vertebrates_CORE_m291_MA0863_1 |
MTF1 |
cluster_125 |
JASPAR_2022_vertebrates_CORE |
14 |
14.15 |
73 |
TTTGCACACGgCAC |
GTGCCGTGTGCAAA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m53_MA0497_1 |
MEF2C |
cluster_30 |
JASPAR_2022_vertebrates_CORE |
16 |
9.72 |
2209 |
rdkcyAAAAATAGmw+ |
+WKCTATTTTTRGMHY |
 |
 |
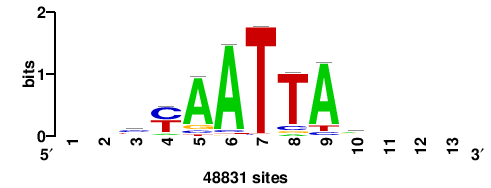
| JASPAR_2022_vertebrates_CORE_m166_MA0717_1 |
RAX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.93 |
48831 |
++yyAATTAr+++ |
+++YTAATTRR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m52_MA0494_1 |
Nr1h3::Rxra |
cluster_58 |
JASPAR_2022_vertebrates_CORE |
24 |
9.43 |
1269 |
++++CMRRGGTYACYYBAGGTCA+ |
+TGaCCtvrrGTrACCyykg++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m435_MA1535_1 |
NR2C1 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
6.88 |
12163 |
++++++++mrrGGTCAy+++ |
+++RTGACCYYK++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m381_MA1419_1 |
IRF4 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
16.09 |
51719 |
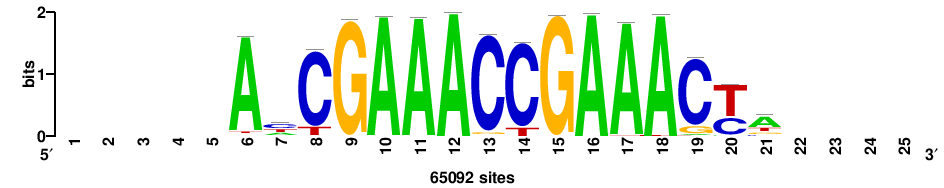
++++++TRGTTTCGGTTTCGR++++ |
++++yCGAAACCGAAACya++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m450_MA1554_1 |
RFX7 |
cluster_25 |
JASPAR_2022_vertebrates_CORE |
17 |
6.47 |
432 |
+RTRGCAACG+++++++ |
+++++++cGTTgCyay+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m37_MA0151_1 |
Arid3a |
cluster_97 |
JASPAR_2022_vertebrates_CORE |
6 |
6.86 |
27 |
ATyAAA |
TTTRAT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m802_MA0799_2 |
RFX4 |
cluster_25 |
JASPAR_2022_vertebrates_CORE |
17 |
14.38 |
1141 |
YSGTTGCYWRGCAACSR |
ysGTTGCywrGCAACsr |
 |
 |
| JASPAR_2022_vertebrates_CORE_m266_MA0839_1 |
CREB3L1 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
11.96 |
1030 |
++TGRTGACGTGGCAY++ |
++rTGCCACGTCAyCa++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m596_MA0102_4 |
CEBPA |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
8.54 |
79003 |
WWATTGTGCAATMW+ |
+wkATTGCACAAtww |
 |
 |
| JASPAR_2022_vertebrates_CORE_m458_MA1562_1 |
SOX14 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
8.78 |
1255 |
+cCGAACAATg++++++++ |
++++++++CATTGTTCGG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m188_MA0736_1 |
GLIS2 |
cluster_46 |
JASPAR_2022_vertebrates_CORE |
16 |
11.69 |
1621 |
+kACCCCCCrCramG+ |
+CKTYGYGGGGGGTM+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m162_MA0713_1 |
PHOX2A |
cluster_38 |
JASPAR_2022_vertebrates_CORE |
16 |
8.36 |
17849 |
++TAAtyyAATTA+++ |
+++TAATTRRATTA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m623_MA0491_2 |
JUND |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.86 |
105945 |
+++wraTGACTCAThh++++ |
++++DDATGAGTCATYW+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m524_MA0912_2 |
HOXD3 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
8.14 |
19015 |
++CTAATTAc+++ |
+++GTAATTAG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m447_MA1550_1 |
PPARD |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
13.96 |
14540 |
++DTGACCTTTGACCYYT+ |
+arrGGTCAAAGGTCAh++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m652_MA0081_2 |
SPIB |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.12 |
26930 |
++RAAAGAGGAAGTGARA++++ |
++++tytCACTTCCTCttty++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m396_MA1480_1 |
DPRX |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
7.24 |
12850 |
SRGATTAKCT++ |
++aGmTAATCys |
 |
 |
| JASPAR_2022_vertebrates_CORE_m629_MA0052_4 |
MEF2A |
cluster_30 |
JASPAR_2022_vertebrates_CORE |
16 |
8.60 |
16162 |
+wkcTAAAAATAGaww |
WWTCTATTTTTAGMW+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m209_MA0773_1 |
MEF2D |
cluster_30 |
JASPAR_2022_vertebrates_CORE |
16 |
12.82 |
6585 |
++dCTAwAAATAGm++ |
++KCTATTTWTAGH++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m814_MA0879_2 |
DLX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.68 |
8937 |
++syAATTAs+++ |
+++STAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m751_MA0136_3 |
Elf5 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
7.96 |
42146 |
++++rgAAGGAAGTrr++++++ |
++++++YYACTTCCTTCY++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m180_MA0728_1 |
Nr2F6 |
cluster_47 |
JASPAR_2022_vertebrates_CORE |
19 |
17.36 |
719 |
+rAGGTCAArAGGTCA+++ |
+++TGACCTYTTGACCTY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m302_MA0881_1 |
Dlx4 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.25 |
15176 |
++syAATTAm+++ |
+++KTAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m363_MA1136_1 |
FOSB::JUNB |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
7.73 |
1001 |
+++++RTGACGTCAY+++++ |
+++++rTGACGTCAy+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m196_MA0088_2 |
ZNF143 |
cluster_63 |
JASPAR_2022_vertebrates_CORE |
21 |
15.29 |
2507 |
++ywCCCAyAATGCAyyG+++ |
+++CRRTGCATTRTGGGWR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m109_MA0131_2 |
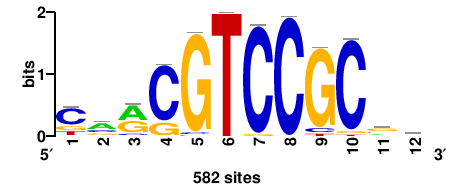
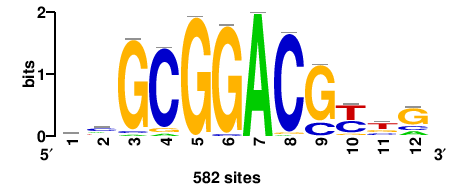
HINFP |
cluster_116 |
JASPAR_2022_vertebrates_CORE |
12 |
9.03 |
582 |
carCGTCCGCgk |
MCGCGGACGYTG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m418_MA1513_1 |
KLF15 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.96 |
11369 |
++scCCCGCCCcs+++++ |
+++++SGGGGCGGGGS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m821_MA1476_2 |
Dlx5 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.94 |
4077 |
wwGCAATTAgmw+ |
+WKCTAATTGCWW |
 |
 |
| JASPAR_2022_vertebrates_CORE_m124_MA0664_1 |
MLXIPL |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
8.82 |
814 |
++++rtCACGTGat++++ |
++++ATCACGTGAY++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m793_MA0742_2 |
KLF12 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
10.05 |
998 |
+++CCCCGCCCC++++++ |
++++++GGGGCGGGG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m462_MA1569_1 |
TFAP2E |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
8.28 |
21373 |
+++ysCCysrGGsr+ |
+YSCCYSRGGSR+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m818_MA1110_2 |
Nr1H4 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
7.46 |
4736 |
++++++++MSTGACCTSB++ |
++vsAGGTCAsk++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m150_MA0695_1 |
ZBTB7C |
cluster_36 |
JASPAR_2022_vertebrates_CORE |
14 |
7.24 |
2132 |
+rcgACCACCgam+ |
+KTCGGTGGTCGY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m289_MA0861_1 |
TP73 |
cluster_50 |
JASPAR_2022_vertebrates_CORE |
18 |
12.14 |
16416 |
KRCRTGTYYRGACATGYH |
drCATGTcyrrACAyGym |
 |
 |
| JASPAR_2022_vertebrates_CORE_m213_MA0776_1 |
MYBL1 |
cluster_48 |
JASPAR_2022_vertebrates_CORE |
17 |
10.59 |
5051 |
+++RCSGTTAACGGT++ |
++ACCGTTAACsgy+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m51_MA0492_1 |
JUND |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
10.41 |
33631 |
+wrwrATGAyGTMATm++++ |
++++KATKACRTCATYWYW+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m390_MA1471_1 |
BARX2 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
7.18 |
52734 |
+++++awwAAymRTTAs++ |
++STAAYKRTTWWT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m464_MA1571_1 |
TGIF2LX |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
9.65 |
3276 |
+TGACAGCTGTCA++ |
++TGACAgcTGTCA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m249_MA0813_1 |
TFAP2B |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
9.95 |
1392 |
+TGCCCYVRGGGCA+ |
+tGCCCybrGGGCa+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m706_MA1967_1 |
TFAP4::FLI1 |
cluster_40 |
JASPAR_2022_vertebrates_CORE |
16 |
14.70 |
1688 |
RWCAGCTGTTTCCGGT |
acCGGAAACAGCTGwy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m501_MA1625_1 |
Stat5b |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
9.08 |
2760 |
++SAYTTCTGGGAAWTR+ |
+yawTTCCcAGAArts++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m430_MA1529_1 |
NHLH2 |
cluster_75 |
JASPAR_2022_vertebrates_CORE |
18 |
12.76 |
26266 |
ggGyMGCAGCTGCGyCmc |
GKGRCGCAGCTGCKRCCC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m348_MA1124_1 |
ZNF24 |
cluster_90 |
JASPAR_2022_vertebrates_CORE |
18 |
12.06 |
23304 |
+++++GAATGAATGAATG |
CATTCATTCATTC+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m699_MA1960_1 |
MGA::EVX1 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
9.76 |
4516 |
++++++++++WMATTAWCACCY |
rGGTGwTAATkw++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m160_MA0711_1 |
OTX1 |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
6.54 |
36255 |
++SGGATTAR++ |
++yTAATCcs++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m336_MA1114_1 |
PBX3 |
cluster_34 |
JASPAR_2022_vertebrates_CORE |
17 |
9.08 |
7021 |
gkbTGAGTGACAGgcrs |
SYGCCTGTCACTCAVMC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m325_MA0852_2 |
FOXK1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
7.95 |
1056 |
++++wrkGTAAACAagsv+ |
+BSCTTGTTTACMYW++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m295_MA0872_1 |
TFAP2A |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
9.77 |
1911 |
+tGCCCysrGGGCa+ |
+TGCCCYSRGGGCA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m65_MA0150_2 |
Nfe2l2 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
11.00 |
726 |
+casmrTGACTCAGCA++++ |
++++TGCTGAGTCAYKSTG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m404_MA1496_1 |
HOXA4 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.62 |
3421 |
++YTAATKRY+++ |
+++rYmATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m137_MA0678_1 |
OLIG2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.98 |
2501 |
++RMCATATGKT+++ |
+++AmCATATGky++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m284_MA0858_1 |
Rarb |
cluster_31 |
JASPAR_2022_vertebrates_CORE |
18 |
17.19 |
769 |
+AGGTCAhmyarAGGTCA |
TGACCTYTRKDTGACCT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m337_MA1115_1 |
POU5F1 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
7.14 |
13114 |
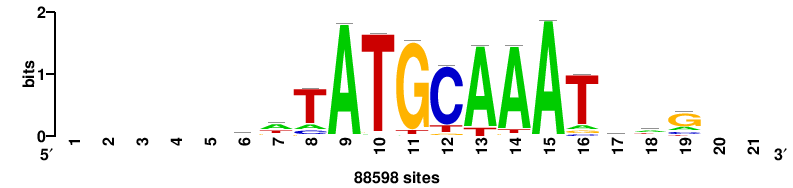
++++HATTTGCATWW++++++ |
++++++wwATGCAAAtd++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m817_MA0909_3 |
Hoxd13 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
8.44 |
5977 |
++TTTTATTGST+++ |
+++asCAATAAAa++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m561_MA1632_1 |
ATF2 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
9.59 |
59394 |
+++wrATGAbGTCAty++++ |
++++RATGACVTCATYW+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m130_MA0048_2 |
NHLH1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
9.14 |
4699 |
++cGCAGCTGCk+++ |
+++MGCAGCTGCG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m440_MA1541_1 |
NR6A1 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
10.83 |
1972 |
gtcAAGkTCAAGktCRw+++ |
+++WYGAMCTTGAMCTTGAC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m727_MA1988_1 |
Atf3 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
7.40 |
54692 |
++++drTGACTCAth+++++ |
+++++DATGAGTCAYH++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m30_MA0135_1 |
Lhx3 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
11.34 |
20 |
+++wAATTAATTAwyy+++ |
+++RRWTAATTAATTW+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m42_MA0259_1 |
ARNT::HIF1A |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
6.75 |
104 |
+++++GCACGTVB+++++ |
+++++vbACGTGc+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m313_MA0895_1 |
HMBOX1 |
cluster_81 |
JASPAR_2022_vertebrates_CORE |
12 |
6.64 |
1642 |
++myTAGTTAms |
SKTAACTARK++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m514_MA0038_2 |
GFI1 |
cluster_55 |
JASPAR_2022_vertebrates_CORE |
12 |
10.70 |
32495 |
cmAATCACdGCa |
TGCHGTGATTKG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m712_MA1973_1 |
ZKSCAN3 |
cluster_137 |
JASPAR_2022_vertebrates_CORE |
20 |
12.37 |
210 |
kkkCCcAGGCTAGCCCasrs |
SYSTGGGCTAGCCTGGGMMM |
 |
 |
| JASPAR_2022_vertebrates_CORE_m580_MA1653_1 |
ZNF148 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.39 |
11199 |
+ysCCCCTCCCcc+++++ |
+++++GGGGGAGGGGSR+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m555_MA1720_1 |
ZNF85 |
cluster_68 |
JASPAR_2022_vertebrates_CORE |
16 |
10.73 |
584 |
amrrAGATTaCwkCAk |
MTGMWGTAATCTYYKT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m301_MA0880_1 |
Dlx3 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.87 |
9548 |
++syAATTAm+++ |
+++KTAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m40_MA0163_1 |
PLAG1 |
cluster_82 |
JASPAR_2022_vertebrates_CORE |
14 |
13.41 |
18 |
GGGGCCcwmGGGGG |
CCCCCKWGGGCCCC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m713_MA1974_1 |
ZNF211 |
cluster_70 |
JASPAR_2022_vertebrates_CORE |
20 |
10.78 |
794 |
+++wwyATATACCAyryw++ |
++WRYRTGGTATATRWW+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m281_MA0855_1 |
RXRB |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
16.02 |
7552 |
+++TGACCTTTGACCYY++ |
++rRGGTCAAAGGTCA+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m586_MA1725_1 |
ZNF189 |
cluster_57 |
JASPAR_2022_vertebrates_CORE |
15 |
8.60 |
15946 |
chtgCTGTTCCht++ |
++ADGGAACAGCADG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m214_MA0777_1 |
MYBL2 |
cluster_48 |
JASPAR_2022_vertebrates_CORE |
17 |
11.63 |
2540 |
++RRCSGTTWAACGGYY |
rrCCGTTwAACsGyy++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m86_MA0622_1 |
Mlxip |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
8.69 |
1000 |
+++++bCACGTGk+++++ |
+++++MCACGTGV+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m224_MA0788_1 |
POU3F3 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
9.36 |
1696 |
+++WAATTWGCATAWW+++++ |
+++++wwTATGCwAATtw+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m439_MA1539_1 |
NR2F6 |
cluster_14 |
JASPAR_2022_vertebrates_CORE |
16 |
14.25 |
12412 |
rAGGTCAbTGACCTy+ |
+RAGGTCAVTGACCTY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m191_MA0739_1 |
Hic1 |
cluster_49 |
JASPAR_2022_vertebrates_CORE |
12 |
6.60 |
15344 |
+rTGCCAaCc++ |
++GGTTGGCAY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m486_MA1600_1 |
ZNF684 |
cluster_2 |
JASPAR_2022_vertebrates_CORE |
19 |
14.48 |
1520 |
+YMAAGGGGTGGACTGTRY |
ryACAGTCCACCCCTTkr+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m405_MA1497_1 |
HOXA6 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.85 |
13284 |
+ggymATTAas++ |
++STTAATKRCC+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m665_MA0609_2 |
CREM |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
8.08 |
17027 |
++GCRGTGACGTCACYGC++ |
++gcrgTGACGTCAcygc++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m601_MA0154_4 |
EBF1 |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
8.95 |
45425 |
WGTCCCCTGGGGAYW |
wrTCCCCAGGGGAcw |
 |
 |
| JASPAR_2022_vertebrates_CORE_m136_MA0677_1 |
Nr2f6 |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
12.60 |
695 |
+++TGACCTTTGACCYY++ |
++rrGGTCAAAGGTCA+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m437_MA1537_1 |
NR2F1 |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
14.06 |
41192 |
+++rrGGTCAAAGGTCAm+ |
+KTGACCTTTGACCYY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m139_MA0683_1 |
POU4F2 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
14.51 |
4552 |
++CTMATTAATTATKCAT+ |
+aTGmATAATTAATkAg++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m680_MA1941_1 |
ETV2::FIGLA |
cluster_40 |
JASPAR_2022_vertebrates_CORE |
16 |
10.91 |
20459 |
rsCGGAArCAGsTGgd |
HCCASCTGYTTCCGSY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m98_MA0641_1 |
ELF4 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
11.27 |
1309 |
+++aACCCGGAAGTr+++++++ |
+++++++YACTTCCGGGTT+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m541_MA0122_3 |
Nkx3-2 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
8.41 |
10220 |
WTTAAGTGKTTWW+++ |
+++wwaamCACTTAaw |
 |
 |
| JASPAR_2022_vertebrates_CORE_m521_MA0902_2 |
HOXB2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.63 |
10302 |
++STAATKRS+++ |
+++symATTAs++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m84_MA0619_1 |
LIN54 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
7.83 |
1001 |
+++++AATTYAAAW+++++++ |
+++++++wTTTrAATt+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m488_MA1603_1 |
Dmrt1 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
9.09 |
18295 |
++GWTACWTTGTAWC++++ |
++++gwtACAAWGTawc++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m77_MA0604_1 |
Atf1 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
7.29 |
1001 |
+++++rTGACGTa+++++++ |
+++++++TACGTCAY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m441_MA1542_1 |
OSR1 |
cluster_56 |
JASPAR_2022_vertebrates_CORE |
15 |
8.61 |
6413 |
++MACRGTAGCR+++ |
+++yGCTACyGTk++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m434_MA1534_1 |
NR1I3 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
7.46 |
67134 |
++++++++AAAGTTCAY+++ |
+++rTGAACTtt++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m78_MA0608_1 |
Creb3l2 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
9.01 |
1000 |
++++KmCACGTGd+++++ |
+++++HCACGTGKM++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m608_MA0477_2 |
FOSL1 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.70 |
57681 |
+++draTGACTCAthy++++ |
++++RDATGAGTCATYH+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m468_MA1576_1 |
THRB |
cluster_58 |
JASPAR_2022_vertebrates_CORE |
24 |
14.99 |
551 |
+++rTGACCTyacrTGACCTyA++ |
++TRAGGTCAYGTRAGGTCAY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m50_MA0488_1 |
JUN |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
10.49 |
20968 |
++awrATGAtGTmAT+++++ |
+++++ATKACATCATYWT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m734_MA1995_1 |
Npas4 |
cluster_20 |
JASPAR_2022_vertebrates_CORE |
14 |
6.27 |
8960 |
++vrTCGTGACws+ |
+SWGTCACGAYB++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m813_MA0878_3 |
CDX1 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
9.60 |
10576 |
++TTTTATKRCC+++ |
+++GGymATAAAA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m729_MA1990_1 |
Gli1 |
cluster_36 |
JASPAR_2022_vertebrates_CORE |
14 |
8.79 |
4003 |
mvrGACCACCCAsr |
YSTGGGTGGTCYBK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m431_MA1530_1 |
NKX6-3 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.31 |
11827 |
+TWTAATGRS+++ |
+++syCATTAwa+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m576_MA1649_1 |
ZBTB12 |
cluster_16 |
JASPAR_2022_vertebrates_CORE |
15 |
7.12 |
4767 |
++RKGTTCYAGRY++ |
++ryCTrGAACmy++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m22_MA0107_1 |
RELA |
cluster_32 |
JASPAR_2022_vertebrates_CORE |
18 |
10.23 |
18 |
+++gGGraTTTCC+++++ |
+++++GGAAATYCCC+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m278_MA0851_1 |
Foxj3 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
9.23 |
101 |
++hraarRTAAACAAwsmc |
GKSWTTGTTTAYYTTYD++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m824_MA1491_2 |
GLI3 |
cluster_46 |
JASPAR_2022_vertebrates_CORE |
16 |
15.54 |
10666 |
rGACCACCCACGwhGh |
DCDWCGTGGGTGGTCY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m788_MA0707_2 |
MNX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.36 |
38982 |
++syrATTAk+++ |
+++MTAATYRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m89_MA0629_1 |
Rhox11 |
cluster_115 |
JASPAR_2022_vertebrates_CORE |
17 |
8.32 |
101 |
amkmyGCTGTwAwrsrw |
WYSYWTWACAGCRKMKT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m719_MA1980_1 |
ZNF418 |
cluster_129 |
JASPAR_2022_vertebrates_CORE |
15 |
12.27 |
999 |
mrGaRGCTAAAAGCA |
TGCTTTTAGCYTCYK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m140_MA0512_2 |
Rxra |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
15.70 |
52026 |
+++grGGTCAAAGGTCA++ |
++TGACCTTTGACCYC+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m552_MA1717_1 |
ZNF784 |
cluster_66 |
JASPAR_2022_vertebrates_CORE |
11 |
8.61 |
12776 |
RCGGTARGTAC |
gtACyTACCgy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m625_MA1107_2 |
KLF9 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.55 |
25579 |
crgCCACACCCAChyc++ |
++GRDGTGGGTGTGGCYG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m27_MA0119_1 |
NFIC::TLX1 |
cluster_28 |
JASPAR_2022_vertebrates_CORE |
21 |
13.63 |
16 |
+++TTGGCWYSSTGCCA++++ |
++++TGGCAssrwGCCAA+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m466_MA1574_1 |
THRB |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
11.06 |
12667 |
+++rrGGTCAAAGGTCRb+ |
+VYGACCTTTGACCYY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m394_MA1478_1 |
DMRTA2 |
cluster_26 |
JASPAR_2022_vertebrates_CORE |
12 |
6.56 |
3681 |
AWTGTAWCAATT |
aattGwTACAwt |
 |
 |
| JASPAR_2022_vertebrates_CORE_m506_MA0605_2 |
ATF3 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
11.96 |
42066 |
++++GRTGACGTCAYC++++ |
++++grTGACGTCAyc++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m490_MA1606_1 |
Foxf1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
7.28 |
59540 |
+++++awrTAAACAaa+++ |
+++TTTGTTTAYWT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m231_MA0600_2 |
RFX2 |
cluster_25 |
JASPAR_2022_vertebrates_CORE |
17 |
14.03 |
15719 |
+sGTTrCCATGGyAACs |
SGTTRCCATGGYAACS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m530_MA0703_2 |
LMX1B |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
9.79 |
5548 |
++TTAATTAAAWT |
awtTTAATTAa++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m684_MA1945_1 |
ETV5::FIGLA |
cluster_40 |
JASPAR_2022_vertebrates_CORE |
16 |
12.20 |
3086 |
CCACCTGYTTCCKCY+ |
+rgmGGAArCAGGTGg |
 |
 |
| JASPAR_2022_vertebrates_CORE_m554_MA1719_1 |
ZNF816 |
cluster_112 |
JASPAR_2022_vertebrates_CORE |
18 |
11.37 |
1053 |
rrrkGGGGACATGcwgGg |
CCCWGCATGTCCCCMYYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m219_MA0781_1 |
PAX9 |
cluster_23 |
JASPAR_2022_vertebrates_CORE |
18 |
15.43 |
5014 |
+sGTCACGCwTSAsTGmm |
KKCASTSAWGCGTGACS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m683_MA1944_1 |
ETV5::DRGX |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.54 |
626 |
+++++rsmGGAAGyAATTA+++ |
+++TAATTRCTTCCKSY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m144_MA0009_2 |
TBXT |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
13.15 |
11810 |
++TmRCACcTAgGTGyGA++++ |
++++TCRCACCTAGGTGYKA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m746_MA0078_2 |
Sox17 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
7.91 |
12287 |
rvagAACAATGGmm+++++ |
+++++KKCCATTGTTCTBY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m598_MA0018_4 |
CREB1 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
8.00 |
79544 |
+++wdrTGATGTCAyh++++ |
++++DRTGACATCAYHW+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m211_MA0774_1 |
MEIS2 |
cluster_34 |
JASPAR_2022_vertebrates_CORE |
17 |
7.60 |
2337 |
++++++tTGACAGs+++ |
+++SCTGTCAA++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m798_MA0765_3 |
ETV5 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.88 |
23301 |
+++++ACCGGAAGTr+++++++ |
+++++++YACTTCCGGT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m754_MA0157_3 |
Foxo3 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
7.01 |
23912 |
+++++wwGTAAACAara++ |
++TYTTGTTTACWW+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m456_MA1560_1 |
SOHLH2 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.83 |
1924 |
++++YGCACGTGCR++++ |
++++ygCACGTGcr++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m198_MA0755_1 |
CUX2 |
cluster_35 |
JASPAR_2022_vertebrates_CORE |
10 |
8.10 |
86361 |
TrATCRATaa |
TTATYGATYA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m633_MA0500_2 |
MYOG |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.15 |
22357 |
+SARCAGCTGYTS++ |
++sarCAGCTGyts+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m717_MA1978_1 |
ZNF354A |
cluster_95 |
JASPAR_2022_vertebrates_CORE |
21 |
8.34 |
999 |
hwAAwtrTAAAtGraywAwtT |
AAWTWRTYCATTTAYAWTTWD |
 |
 |
| JASPAR_2022_vertebrates_CORE_m120_MA0660_1 |
MEF2B |
cluster_30 |
JASPAR_2022_vertebrates_CORE |
16 |
12.73 |
3872 |
++rCTAwAAATAGm++ |
++KCTATTTWTAGY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m433_MA1533_1 |
NR1I2 |
cluster_61 |
JASPAR_2022_vertebrates_CORE |
17 |
10.23 |
241 |
GRGTTCRTYSRGKTCRY |
ryGAMCysraYGAACyc |
 |
 |
| JASPAR_2022_vertebrates_CORE_m67_MA0144_2 |
STAT3 |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
9.04 |
21620 |
++++yTKCykGGAAr+++ |
+++YTTCCMRGMAR++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m96_MA0638_1 |
CREB3 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
10.73 |
539 |
++YGRTGACGTGKCAY++ |
++rtGmCACGTCAycr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m212_MA0775_1 |
MEIS3 |
cluster_34 |
JASPAR_2022_vertebrates_CORE |
17 |
6.25 |
12838 |
++++++wTGACAgs+++ |
+++SCTGTCAW++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m323_MA0911_1 |
Hoxa11 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
7.93 |
1258 |
+AWTTTWAYGACY |
rgTCRTwAAawt+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m251_MA0815_1 |
TFAP2C |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
10.04 |
3137 |
+tGCCCysrGGGCa+ |
+TGCCCYSRGGGCA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m789_MA0708_2 |
MSX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.89 |
9310 |
++sYAATTAr+++ |
+++YTAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m726_MA1987_1 |
ZNF701 |
cluster_100 |
JASPAR_2022_vertebrates_CORE |
21 |
11.65 |
999 |
GAGmAsyaarGGGGGrArdkg |
CMHYTYCCCCCYTTRSTKCTC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m635_MA0089_2 |
MAFG::NFE2L1 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
12.30 |
3356 |
++avmATGACTCAGCAaw++ |
++WTTGCTGAGTCATKBT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m19_MA0091_1 |
TAL1::TCF3 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
9.75 |
44 |
mgAmCAKMTGkT+++ |
+++AMCAKMTGKTCK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m718_MA1979_1 |
ZNF416 |
cluster_49 |
JASPAR_2022_vertebrates_CORE |
12 |
8.98 |
345 |
+RTGCCCAGATA |
TaTCTGGGCay+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m267_MA0840_1 |
Creb5 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
11.31 |
4641 |
++++RRTGACGTCATY++++ |
++++rATGACGTCAyy++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m742_MA0037_4 |
Gata3 |
cluster_11 |
JASPAR_2022_vertebrates_CORE |
15 |
7.22 |
24868 |
DVAGATAAGAWH+++ |
+++dwtCTTATCTbh |
 |
 |
| JASPAR_2022_vertebrates_CORE_m686_MA1947_1 |
ETV5::FOXO1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.47 |
434 |
+rtmAACAGGAwrt++++++++ |
++++++++AYWTCCTGTTKAY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m833_MA1566_2 |
TBX3 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
8.91 |
11289 |
+++++++++rAGGTGTGAAm++ |
++KTTCACACCTY+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m841_MA1540_2 |
NR5A1 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
9.52 |
4697 |
+rkragtTCAAGGtCAgc++ |
++GCTGACCTTGAACTYMY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m142_MA0687_1 |
SPIC |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.17 |
367 |
+aaAAWsmGGAAGTa+++++++ |
+++++++TACTTCCKSWTTTT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m346_MA0750_2 |
ZBTB7A |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
7.94 |
14641 |
++++rsCCGGAAGtgss+++++ |
+++++SSCACTTCCGGSY++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m705_MA1966_1 |
TFAP4::ETV1 |
cluster_40 |
JASPAR_2022_vertebrates_CORE |
16 |
10.38 |
4498 |
rsCGGAwgCAGsTGgk |
MCCASCTGCWTCCGSY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m116_MA0655_1 |
JDP2 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
10.47 |
2065 |
+++++ATGASTCAT++++++ |
++++++ATGASTCAT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m322_MA0908_1 |
HOXD11 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
8.57 |
412 |
++WTTTTACGAY+ |
+rTCGTAAAaw++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m816_MA0885_2 |
Dlx2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.79 |
7437 |
awGCAATTAGmw+ |
+WKCTAATTGCWT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m190_MA0738_1 |
HIC2 |
cluster_49 |
JASPAR_2022_vertebrates_CORE |
12 |
5.86 |
5890 |
+rTGCCMrsy++ |
++RSYKGGCAY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m407_MA1500_1 |
HOXB6 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.54 |
11660 |
++tTAATkrc+++ |
+++GYMATTAA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m565_MA1637_1 |
EBF3 |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
8.48 |
22062 |
++kyCCCAAGGGAvw |
WBTCCCTTGGGRM++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m297_MA0873_1 |
HOXD12 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
9.83 |
2251 |
+KTTTTACGACY+ |
+rgTCGTAAAAm+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m831_MA1547_2 |
PITX2 |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
8.37 |
8687 |
++yTAATCCy++ |
++RGGATTAR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m823_MA1487_2 |
FOXE1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
10.43 |
4583 |
++syTAARyAAACAaw+++ |
+++WTTGTTTRYTTARS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m345_MA1122_1 |
TFDP1 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
7.73 |
2007 |
++++gsGCGGGAAss+++ |
+++SSTTCCCGCSC++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m93_MA0007_3 |
Ar |
cluster_19 |
JASPAR_2022_vertebrates_CORE |
17 |
12.94 |
3851 |
RRGWACAYCRTGTWCYY |
rRGwACAygrTGTwCyy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m728_MA1989_1 |
Bcl11B |
cluster_60 |
JASPAR_2022_vertebrates_CORE |
14 |
7.46 |
39791 |
KTYTTGTGGTTTTT |
aaaAACCACAaram |
 |
 |
| JASPAR_2022_vertebrates_CORE_m787_MA0697_2 |
Zic3 |
cluster_56 |
JASPAR_2022_vertebrates_CORE |
15 |
7.96 |
26174 |
++caCAGCAGGaggs |
SCCTCCTGCTGTG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m803_MA0818_2 |
BHLHE22 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
9.29 |
10475 |
++RMCATATGKT+++ |
+++AmCATATGky++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m772_MA0611_2 |
Dux |
cluster_68 |
JASPAR_2022_vertebrates_CORE |
16 |
11.47 |
3067 |
WWWGTGATTVAATCWR |
ywGATTbAATCAcwww |
 |
 |
| JASPAR_2022_vertebrates_CORE_m720_MA1981_1 |
ZNF530 |
cluster_99 |
JASPAR_2022_vertebrates_CORE |
18 |
12.23 |
999 |
GmArGGmrAGGGsCrgsv |
BSCYGSCCCTYKCCYTKC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m364_MA1137_1 |
FOSL1::JUNB |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
6.35 |
1001 |
+++ykrTGAcTMAtmc++++ |
++++GKATKAGTCAYMR+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m341_MA1119_1 |
SIX2 |
cluster_33 |
JASPAR_2022_vertebrates_CORE |
16 |
9.14 |
5453 |
RKATCAGGTTTCAGWW |
wwcTGaAACCTGAtmy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m614_MA0035_4 |
GATA1 |
cluster_11 |
JASPAR_2022_vertebrates_CORE |
15 |
7.00 |
71828 |
WWAGATWAGAW++++ |
++++wtCTWATCTww |
 |
 |
| JASPAR_2022_vertebrates_CORE_m332_MA0161_2 |
NFIC |
cluster_49 |
JASPAR_2022_vertebrates_CORE |
12 |
7.34 |
11607 |
TSTGCCAAGWA+ |
+twCTTGGCAsa |
 |
 |
| JASPAR_2022_vertebrates_CORE_m504_MA1629_1 |
Zic2 |
cluster_56 |
JASPAR_2022_vertebrates_CORE |
15 |
9.19 |
11919 |
CYYCCTGCTGTGMK+ |
+mkCaCAGCAGGrrg |
 |
 |
| JASPAR_2022_vertebrates_CORE_m757_MA0467_2 |
Crx |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
7.57 |
4333 |
+BYTAATCCYC+ |
+grGGATTArv+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m300_MA0876_1 |
BSX |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.90 |
10170 |
++BTAATKRS+++ |
+++symATTAv++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m566_MA1638_1 |
HAND2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.82 |
30283 |
++RVCATCTGKT+++ |
+++amCAGATGby++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m317_MA0899_1 |
HOXA10 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
6.34 |
14876 |
++rgymATWAAaa++ |
++TTTTWATKRCY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m415_MA1508_1 |
IKZF1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
7.86 |
10918 |
++rrAACAGGAAry++++++++ |
++++++++RYTTCCTGTTYY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m377_MA1151_1 |
RORC |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
6.92 |
1001 |
++++dwAwgTrGGtCA++++ |
++++TGACCYACWTWH++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m492_MA1608_1 |
Isl1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.97 |
5637 |
+KRCTAATGGMT+ |
+akCCATTAgym+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m631_MA1108_2 |
MXI1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.52 |
22048 |
++rvCACATGkc+++ |
+++GMCATGTGBY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m549_MA1714_1 |
ZNF675 |
cluster_107 |
JASPAR_2022_vertebrates_CORE |
21 |
12.05 |
999 |
rgGmyyarGAGGmcAaAATGw |
WCATTTTGKCCTCYTRRKCCY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m63_MA0076_2 |
ELK4 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.07 |
3427 |
+++++RCCGGAAGYGS++++++ |
++++++scrCTTCCGGy+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m736_MA1997_1 |
Olig2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.19 |
61142 |
++rvCAGCTGby+++ |
+++RVCAGCTGBY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m193_MA0747_1 |
SP8 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
8.41 |
268 |
++rmCACgCCCmCy++++ |
++++RGKGGGCGTGKY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m639_MA1112_2 |
NR4A1 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
8.35 |
15927 |
++++++TBTGACCTTTWM++ |
++kwAAAGGTCAva++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m443_MA1545_1 |
OVOL2 |
cluster_48 |
JASPAR_2022_vertebrates_CORE |
17 |
8.70 |
18908 |
+rwaCCGTTAydys+++ |
+++SRHRTAACGGTWY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m795_MA0756_2 |
ONECUT2 |
cluster_35 |
JASPAR_2022_vertebrates_CORE |
10 |
8.77 |
11001 |
TATYGATCG+ |
+cGATCrATA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m282_MA0856_1 |
RXRG |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
15.48 |
61684 |
+++grGGTCAAAGGTCA++ |
++TGACCTTTGACCYC+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m547_MA1712_1 |
ZNF454 |
cluster_123 |
JASPAR_2022_vertebrates_CORE |
18 |
10.45 |
999 |
aGGCTCssGGcCCykvkG |
CMBMRGGGCCSSGAGCCT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m307_MA0888_1 |
EVX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.42 |
29598 |
+svTaATTAgc++ |
++GCTAATTABS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m697_MA1958_1 |
HOXD12::ELK1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
14.72 |
665 |
++++TTTTACRACTTCCGSYY+ |
+rrsCGGAAGTyGTAAAa++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m478_MA1588_1 |
ZNF136 |
cluster_72 |
JASPAR_2022_vertebrates_CORE |
15 |
12.41 |
3201 |
rwATTCTtGGTTGrc |
GYCAACCAAGAATWY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m559_MA1724_1 |
Rfx6 |
cluster_25 |
JASPAR_2022_vertebrates_CORE |
17 |
8.20 |
8550 |
YKGTTGCTAGGAA++++ |
++++ttCCTAGCAACmr |
 |
 |
| JASPAR_2022_vertebrates_CORE_m525_MA0910_2 |
HOXD8 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.85 |
6682 |
++KTAATKRC+++ |
+++gymATTAm++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m384_MA1463_1 |
ARGFX |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
8.35 |
25318 |
+kcTAATTAra++ |
++TYTAATTAGM+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m664_MA0528_2 |
ZNF263 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
8.10 |
56343 |
++CSTCCTCCCCSC++++ |
++++gsgGGGAGGAsg++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m482_MA1594_1 |
ZNF382 |
cluster_121 |
JASPAR_2022_vertebrates_CORE |
24 |
18.17 |
4252 |
AgGrGAyATyACyACmGAyCCyAC |
GTRGGRTCKGTRGTRATRTCYCCT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m659_MA0003_4 |
TFAP2A |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
8.08 |
15968 |
+vytgCCTCAGGsma |
TKSCCTGAGGCARB+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m454_MA1558_1 |
SNAI1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.67 |
30930 |
++YRCACCTGYH+++ |
+++drCAGGTGyr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m199_MA0757_1 |
ONECUT3 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
11.81 |
7730 |
aaAAAATCrATAaw+++++ |
+++++WTTATYGATTTTTT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m491_MA1607_1 |
Foxl2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.03 |
18101 |
++rwwaTGTAAACAwr+++ |
+++YWTGTTTACATWWY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m334_MA1111_1 |
NR2F2 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
7.78 |
2204 |
+++++++yaAAGGTCAar++ |
++YTTGACCTTTR+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m365_MA1138_1 |
FOSL2::JUNB |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.97 |
1001 |
++++krTGAsTCAy++++++ |
++++++RTGASTCAYM++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m108_MA0649_1 |
HEY2 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
8.30 |
2107 |
++++GrCACGTGyc++++ |
++++GRCACGTGYC++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m762_MA0505_2 |
Nr5A2 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
9.10 |
30542 |
++++AGKTCAAGGTCAB+++ |
+++vTGaCCTTGAmct++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m584_MA1657_1 |
ZNF652 |
cluster_81 |
JASPAR_2022_vertebrates_CORE |
12 |
8.81 |
12373 |
kaAAGrGTTAAw |
WTTAACYCTTTM |
 |
 |
| JASPAR_2022_vertebrates_CORE_m276_MA0849_1 |
FOXO6 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.31 |
8470 |
+++++++GTAAACA+++++ |
+++++TGTTTAC+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m216_MA0778_1 |
NFKB2 |
cluster_32 |
JASPAR_2022_vertebrates_CORE |
18 |
11.43 |
16790 |
++aGGGGAwTCCCCy+++ |
+++RGGGGAWTCCCCT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m366_MA1139_1 |
FOSL2::JUNB |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
9.36 |
1000 |
++++krTGACGTCAym++++ |
++++KRTGACGTCAYM++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m627_MA0495_3 |
MAFF |
cluster_51 |
JASPAR_2022_vertebrates_CORE |
16 |
9.19 |
37746 |
WWWAAAATGCTGACWY |
rwGTCAGCAtTTtwww |
 |
 |
| JASPAR_2022_vertebrates_CORE_m606_MA0592_3 |
ESRRA |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
9.07 |
25995 |
+++++kytCAAGGTCAya++ |
++TRTGACCTTGARM+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m197_MA0752_1 |
ZNF410 |
cluster_88 |
JASPAR_2022_vertebrates_CORE |
17 |
17.00 |
4052 |
tmCATCCCATAATAhtc |
GADTATTATGGGATGKA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m238_MA0801_1 |
MGA |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
7.51 |
44673 |
++++++++++TCACACCT++++ |
++++AGGTGTGA++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m408_MA1501_1 |
HOXB7 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.42 |
1238 |
+gGyAATTArs++ |
++SYTAATTRCC+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m355_MA1128_1 |
FOSL1::JUN |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.49 |
1001 |
+++kkrTGACTCAtmm++++ |
++++KKATGAGTCAYMM+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m799_MA0768_2 |
Lef1 |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
7.53 |
1081 |
+++RASATCAAAGGMAW++ |
++wtkCCTTTGATsty+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m495_MA1618_1 |
Ptf1a |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.53 |
14441 |
DMAACATCTGTKH++ |
++dmaCAGATGttkh |
 |
 |
| JASPAR_2022_vertebrates_CORE_m100_MA0643_1 |
Esrrg |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
9.11 |
27172 |
+++++++tsAAGGTCAw+++ |
+++WTGACCTTSA+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m791_MA0734_3 |
Gli2 |
cluster_36 |
JASPAR_2022_vertebrates_CORE |
14 |
9.52 |
2492 |
+vrGACCACCCAsr |
YSTGGGTGGTCYB+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m329_MA0100_3 |
MYB |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.81 |
1527 |
++KDCAGTTGVK+++ |
+++mbCAACTGhm++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m262_MA0827_1 |
OLIG3 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.93 |
3320 |
++RVCATATGKT+++ |
+++AmCATATGby++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m35_MA0002_2 |
Runx1 |
cluster_60 |
JASPAR_2022_vertebrates_CORE |
14 |
7.21 |
2000 |
+ktyTGTGGttw++ |
++WAACCACARAM+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m230_MA0794_1 |
PROX1 |
cluster_87 |
JASPAR_2022_vertebrates_CORE |
12 |
11.74 |
4064 |
yAAGACGyCTTa |
TAAGRCGTCTTR |
 |
 |
| JASPAR_2022_vertebrates_CORE_m170_MA0721_1 |
UNCX |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.88 |
46677 |
++yyAATTAr+++ |
+++YTAATTRR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m248_MA0812_1 |
TFAP2B |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
8.71 |
4790 |
+++mGCCysAGGCd+ |
+HGCCTSRGGCK+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m111_MA0153_2 |
HNF1B |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
11.26 |
40813 |
+++++RTTAATSATTAAC+ |
+GTTAATsATTAAy+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m414_MA1507_1 |
HOXD4 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.80 |
4589 |
++YTAATKRY+++ |
+++rYmATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m603_MA0598_3 |
EHF |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.42 |
15368 |
++RAAACAGGAAGTGRS+++++ |
+++++sycACTTCCTgttty++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m830_MA1532_2 |
NR1D2 |
cluster_47 |
JASPAR_2022_vertebrates_CORE |
19 |
14.63 |
86217 |
+TrGGTyAGTrGGTCA+++ |
+++TGACCYACTRACCYA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m688_MA1949_1 |
FLI1::DRGX |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.92 |
591 |
+++++ATAATTRCWTCCGGT++ |
++aCCGGAwGyAATTAt+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m460_MA1565_1 |
TBX18 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
7.82 |
1200 |
++++++++YTTCACACCTYY++ |
++rrAGGTGTGAar++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m432_MA1531_1 |
NR1D1 |
cluster_47 |
JASPAR_2022_vertebrates_CORE |
19 |
13.17 |
10840 |
++rGGTyAgTrGGTCAr++ |
++YTGACCYACTRACCY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m135_MA0676_1 |
Nr2e1 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
7.88 |
2024 |
++++++++TTGACTTTT+++ |
+++aaAAGTCAA++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m779_MA0651_2 |
HOXC11 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
11.37 |
17709 |
DTTTTACGACYT+ |
+arGTCGTAAAAh |
 |
 |
| JASPAR_2022_vertebrates_CORE_m780_MA0659_3 |
Mafg |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
10.25 |
20118 |
++rwGmTGACTCAGCag+++ |
+++CTGCTGAGTCAKCWY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m617_MA1105_2 |
GRHL2 |
cluster_33 |
JASPAR_2022_vertebrates_CORE |
16 |
7.82 |
41878 |
arAACAGGTTtk++++ |
++++MAAACCTGTTYT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m829_MA1525_2 |
NFATC4 |
cluster_44 |
JASPAR_2022_vertebrates_CORE |
17 |
8.26 |
15974 |
++++aayGGAAAmw+++ |
+++WKTTTCCRTT++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m342_MA0442_2 |
SOX10 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
7.77 |
2055 |
++rraACAAAGvm++++++ |
++++++KBCTTTGTTYY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m657_MA0090_3 |
TEAD1 |
cluster_2 |
JASPAR_2022_vertebrates_CORE |
19 |
8.71 |
70398 |
+hyACATTCCAGsm+++++ |
+++++KSCTGGAATGTRD+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m97_MA0639_1 |
DBP |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
9.52 |
10298 |
+TRTTAYRTAAYM++ |
++krTTAYRTAAya+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m261_MA0826_1 |
OLIG1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
10.16 |
14950 |
++AMCATATGKT+++ |
+++AmCATATGkT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m620_MA0484_2 |
HNF4G |
cluster_2 |
JASPAR_2022_vertebrates_CORE |
19 |
8.53 |
18222 |
kyCAAAGTCCArr++++++ |
++++++YYTGGACTTTGRM |
 |
 |
| JASPAR_2022_vertebrates_CORE_m246_MA0810_1 |
TFAP2A |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
7.46 |
6897 |
++TGCCYBRGGGSR+ |
+ysCCCyvrGGCa++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m732_MA1993_1 |
Neurod2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.42 |
54024 |
++rvCAGCTGby+++ |
+++RVCAGCTGBY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m305_MA0886_1 |
EMX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.13 |
44114 |
+rsTAATTAgs++ |
++SCTAATTASY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m173_MA0725_1 |
VSX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.60 |
19111 |
++yTAATTAb+++ |
+++VTAATTAR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m510_MA0469_3 |
E2F3 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
14.62 |
35365 |
RWTTTGGCGCCAAAWY++ |
++rWTTTGGCGCCAAAWy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m745_MA0067_2 |
PAX2 |
cluster_23 |
JASPAR_2022_vertebrates_CORE |
18 |
11.92 |
4693 |
YCGTCACGCWYSAYTRV+ |
+bYArTsrwGCGTGACGr |
 |
 |
| JASPAR_2022_vertebrates_CORE_m481_MA1593_1 |
ZNF317 |
cluster_56 |
JASPAR_2022_vertebrates_CORE |
15 |
8.24 |
11463 |
+WRTCTGCTGTYW++ |
++wrACAGCAGAyw+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m469_MA1577_1 |
TLX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.35 |
18502 |
++yyAATTAr+++ |
+++YTAATTRR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m527_MA1140_2 |
JUNB |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
12.34 |
333 |
++++GRTGACGTCATC++++ |
++++gATGACGTCAyc++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m499_MA1623_1 |
Stat2 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
9.14 |
3959 |
+++++++rgAAACAGAAAsw+++++ |
+++++WSTTTCTGTTTCY+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m88_MA0628_1 |
POU6F1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.00 |
1001 |
+AYTAATTART++ |
++ayTAATTArt+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m339_MA1117_1 |
RELB |
cluster_32 |
JASPAR_2022_vertebrates_CORE |
18 |
6.99 |
3290 |
+++++gaATTCCCCks++ |
++SMGGGGAATTC+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m675_MA1936_1 |
ERF::FOXO1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.18 |
627 |
+rwmAACAGGAArks+++++++ |
+++++++SMYTTCCTGTTKWY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m422_MA1517_1 |
KLF6 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
7.95 |
3312 |
+srcCACRCCCm++++++ |
++++++KGGGYGTGGYS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m56_MA0515_1 |
Sox6 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
9.15 |
249 |
++RRRACAAWGG+++++++ |
+++++++cCwTTGTyyy++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m350_MA1154_1 |
ZNF282 |
cluster_89 |
JASPAR_2022_vertebrates_CORE |
17 |
14.53 |
445 |
cTTTCCCmyAACACkas |
STMGTGTTRKGGGAAAG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m733_MA1994_1 |
Nkx2-1 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
7.82 |
16242 |
+++DWKYTCAAGTGBY |
rvCACTTGArmwh+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m800_MA0784_2 |
POU1F1 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
13.91 |
5213 |
avCTmATTwGCATAwt+++++ |
+++++AWTATGCWAATKAGBT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m539_MA0871_2 |
TFEC |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
10.03 |
13032 |
+++GGTCACGTGSK++++ |
++++msCACGTGACc+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m588_MA1727_1 |
ZNF417 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
9.82 |
999 |
+++KTGGCGCCCARYVYK |
mrbrytgGGCGCCAm+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m403_MA1495_1 |
HOXA1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.85 |
10981 |
++sTAATtAs+++ |
+++STAATTAS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m618_MA0043_3 |
HLF |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
9.36 |
24424 |
wdrTTATGYAAymw+ |
+WKRTTRCATAAYHW |
 |
 |
| JASPAR_2022_vertebrates_CORE_m34_MA0138_2 |
REST |
cluster_134 |
JASPAR_2022_vertebrates_CORE |
21 |
16.04 |
1606 |
tTCAGcACCatGGACAGckcC |
GGMGCTGTCCATGGTGCTGAA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m292_MA0024_3 |
E2F1 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
9.04 |
1025 |
++TTTGGCGCCAAA++++ |
++++tttGGCGCCaaa++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m320_MA0906_1 |
HOXC12 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
9.98 |
6266 |
+rGTCGTAAAAh+ |
+DTTTTACGACY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m39_MA0159_1 |
RARA::RXRA |
cluster_31 |
JASPAR_2022_vertebrates_CORE |
18 |
11.09 |
23 |
+TGAMCTSYCSRTGAMCY |
rGkTCAysgrsAGKTCA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m112_MA0486_2 |
HSF1 |
cluster_16 |
JASPAR_2022_vertebrates_CORE |
15 |
12.99 |
2266 |
++TTCTaGAAyrTTC |
GAAYRTTCTAGAA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m335_MA0014_3 |
PAX5 |
cluster_20 |
JASPAR_2022_vertebrates_CORE |
14 |
7.68 |
1412 |
++rrGCGTGACCmb |
VKGGTCACGCYY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m353_MA1126_1 |
FOS::JUN |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
7.19 |
1001 |
RAMKKATGACGTCAYH++++ |
++++drTgACGTCAtmmkty |
 |
 |
| JASPAR_2022_vertebrates_CORE_m417_MA1512_1 |
KLF11 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.19 |
11118 |
++gcCACGCCCmC+++++ |
+++++GKGGGCGTGGC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m33_MA0149_1 |
EWSR1-FLI1 |
cluster_90 |
JASPAR_2022_vertebrates_CORE |
18 |
22.78 |
105 |
GGAAGGAAGGAAGGAAGG |
CCTTCCTTCCTTCCTTCC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m755_MA0160_2 |
NR4A2 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
8.09 |
623 |
++++++ktAAAGGTCA++++ |
++++TGACCTTTAM++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m451_MA1555_1 |
RXRB |
cluster_20 |
JASPAR_2022_vertebrates_CORE |
14 |
11.31 |
2146 |
rrGGTCrTGACCyy |
RRGGTCAYGACCYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m17_MA0077_1 |
SOX9 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
6.29 |
76 |
+++RAACAATRG+++++++ |
+++++++CyATTGTTy+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m714_MA1975_1 |
ZNF214 |
cluster_118 |
JASPAR_2022_vertebrates_CORE |
15 |
9.61 |
999 |
rttCATcAAkgtCCT |
AGGACMTTGATGAAY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m815_MA0884_2 |
DUXA |
cluster_38 |
JASPAR_2022_vertebrates_CORE |
16 |
10.65 |
10537 |
+YTRAYTWARTYAR++ |
++yTrAyTwArTyAr+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m232_MA0795_1 |
SMAD3 |
cluster_15 |
JASPAR_2022_vertebrates_CORE |
12 |
10.82 |
4999 |
yGTCTAGACA++ |
++TGTCTAGACR |
 |
 |
| JASPAR_2022_vertebrates_CORE_m245_MA0808_1 |
TEAD3 |
cluster_2 |
JASPAR_2022_vertebrates_CORE |
19 |
7.98 |
44800 |
+++WGGAATGY++++++++ |
++++++++rCATTCCw+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m723_MA1984_1 |
ZNF667 |
cluster_122 |
JASPAR_2022_vertebrates_CORE |
12 |
8.71 |
999 |
kTAaRAGCTCAr |
YTGAGCTYTTAM |
 |
 |
| JASPAR_2022_vertebrates_CORE_m327_MA1106_1 |
HIF1A |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.17 |
980 |
+++SSGCACGTVC+++++ |
+++++gbACGTGCss+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m609_MA0148_4 |
FOXA1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
7.91 |
322803 |
+++++awGTAAACATdw++ |
++WHATGTTTACWT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m215_MA0105_4 |
NFKB1 |
cluster_32 |
JASPAR_2022_vertebrates_CORE |
18 |
14.22 |
19717 |
++AGGGGAWTCCCCT+++ |
+++AGGGGAwTCCCCt++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m570_MA1642_1 |
NEUROG2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.78 |
34391 |
+BRCCATCTGTYYY+ |
+rrraCAGATGGyv+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m155_MA0705_1 |
Lhx8 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.13 |
7263 |
++YYRATTRG+++ |
+++cYAATYRr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m446_MA1549_1 |
POU6F1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.47 |
9847 |
ASCTCATTAY+++ |
+++rTAATGAGst |
 |
 |
| JASPAR_2022_vertebrates_CORE_m167_MA0718_1 |
RAX |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.61 |
2651 |
+ryyAATTAry++ |
++RYTAATTRRY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m379_MA1153_1 |
Smad4 |
cluster_15 |
JASPAR_2022_vertebrates_CORE |
12 |
8.13 |
1000 |
++KCYAGACR++ |
++yGTCTrGm++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m13_MA0071_1 |
RORA |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
9.14 |
25 |
++++++wwcwAGGTCA++++ |
++++TGACCTWGWW++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m677_MA1938_1 |
ERF::NHLH1 |
cluster_96 |
JASPAR_2022_vertebrates_CORE |
17 |
14.27 |
367 |
aGCAGCTGCCGGAWrym |
KRYWTCCGGCAGCTGCT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m356_MA1129_1 |
FOSL1::JUN |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
7.11 |
1001 |
+++++RTGACGTCAY+++++ |
+++++rTgACGTcAy+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m244_MA0807_1 |
TBX5 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
5.88 |
3461 |
++++++++++AGGTGTkA++++ |
++++TMACACCT++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m674_MA1935_1 |
ERF::FOXI1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
7.16 |
1576 |
+rtaaACmGGAArtr+++++++ |
+++++++YAYTTCCKGTTTAY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m70_MA0593_1 |
FOXP2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
9.57 |
766 |
+++++awGTAAACAra+++ |
+++TYTGTTTACWT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m797_MA0764_3 |
ETV4 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.03 |
25381 |
+++++YACTTCCGGT+++++++ |
+++++++aCCGGAAGtr+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m640_MA0679_2 |
ONECUT1 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
9.87 |
62219 |
aaaaAATCAATAawww+++ |
+++WWWTTATTGATTTTTT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m485_MA1599_1 |
ZNF682 |
cluster_32 |
JASPAR_2022_vertebrates_CORE |
18 |
10.35 |
1058 |
AYWGGGGCTTRGCCYG++ |
++crGGCyAAGCCCCwrt |
 |
 |
| JASPAR_2022_vertebrates_CORE_m269_MA0841_1 |
NFE2 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
11.71 |
21530 |
++++vATGACTCATs+++++ |
+++++SATGAGTCATB++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m241_MA0804_1 |
TBX19 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
12.12 |
61661 |
wtTmrCACcTAgGTGygAaa++ |
++TTTCRCACCTAGGTGYKAAW |
 |
 |
| JASPAR_2022_vertebrates_CORE_m507_MA0462_2 |
BATF::JUN |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
7.77 |
38154 |
++++waTGACTCAth+++++ |
+++++DATGAGTCATW++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m634_MA0056_2 |
MZF1 |
cluster_32 |
JASPAR_2022_vertebrates_CORE |
18 |
7.81 |
8242 |
+++++mwAATCCCCAchy |
RDGTGGGGATTWK+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m310_MA0891_1 |
GSC2 |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
5.50 |
3059 |
+cyTAATCcsh+ |
+DSGGATTARG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m263_MA0832_1 |
Tcf21 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
12.55 |
199 |
ryAACAGCTGTTry+ |
+RYAACAGCTGTTRY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m147_MA0691_1 |
TFAP4 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
9.81 |
4837 |
++AwCAGCTGwT+++ |
+++AWCAGCTGWT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m308_MA0889_1 |
GBX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.52 |
28843 |
+rsyAATTAgc++ |
++GCTAATTRSY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m43_MA0145_2 |
Tfcp2l1 |
cluster_119 |
JASPAR_2022_vertebrates_CORE |
14 |
8.09 |
4092 |
CCrGyyyrarCCrG |
CYGGYTYRRRCYGG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m125_MA0665_1 |
MSC |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
10.30 |
94 |
++AACAGCTGTT+++ |
+++AACAGCTGTT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m590_MA1729_1 |
ZNF680 |
cluster_73 |
JASPAR_2022_vertebrates_CORE |
15 |
11.51 |
5766 |
rwCCAAGAAGAATda |
THATTCTTCTTGGWY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m258_MA0823_1 |
HEY1 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
8.50 |
3822 |
++++GRCACGTGYC++++ |
++++grCACGTGyc++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m426_MA1522_1 |
MAZ |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.68 |
15063 |
+GGGGAGGGGSS++++++ |
++++++ssCCCCTCCcc+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m810_MA0865_2 |
E2F8 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
7.77 |
4054 |
+TWWTGGCGGGAA+++++ |
+++++tTCCCGCcAwwa+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m572_MA1644_1 |
NFYC |
cluster_17 |
JASPAR_2022_vertebrates_CORE |
13 |
7.30 |
13616 |
++rrCCAATCAsa |
TSTGATTGGYY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m288_MA0106_3 |
TP53 |
cluster_50 |
JASPAR_2022_vertebrates_CORE |
18 |
14.32 |
23732 |
RACATGYCCGGRCATGTY |
rACATGyCcgGrCATGTy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m581_MA1654_1 |
ZNF16 |
cluster_71 |
JASPAR_2022_vertebrates_CORE |
23 |
17.60 |
833 |
AvTrGGGAGCCATgGArGGtTyt |
ARAACCYTCCATGGCTCCCYABT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m696_MA1957_1 |
HOXB2::ELK1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
11.92 |
1825 |
+++++rcCGGAAGTmrTTA+++ |
+++TAAYKACTTCCGGY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m163_MA0714_1 |
PITX3 |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
6.22 |
8661 |
+vyTAATCCc++ |
++GGGATTARB+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m145_MA0688_1 |
TBX2 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
7.11 |
6189 |
+++++++++TTTCACACCTW++ |
++wAGGTGTgAaa+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m457_MA1561_1 |
SOX12 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
9.24 |
1368 |
aCCGAACAATg++++++++ |
++++++++CATTGTTCGGT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m512_MA0612_2 |
EMX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.33 |
10779 |
++STAATTAS+++ |
+++sTAATTAs++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m91_MA0631_1 |
Six3 |
cluster_105 |
JASPAR_2022_vertebrates_CORE |
17 |
10.45 |
101 |
rAtrGGGTATCAywwwt |
AWWWRTGATACCCYATY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m227_MA0791_1 |
POU4F3 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
9.57 |
7674 |
++CTCATTAATWATKCAT+ |
+aTgmATwATTaATgag++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m71_MA0595_1 |
SREBF1 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
9.37 |
58 |
+++++++++RTGGGGTGAY+++ |
+++rTCACcCCAy+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m69_MA0591_1 |
Bach1::Mafk |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
12.70 |
114 |
++rssrTGACTCAGCAs+++ |
+++STGCTGAGTCAYSSY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m9_MA0059_1 |
MAX::MYC |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
9.87 |
21 |
+++rAsCACGTGGt++++ |
++++ACCACGTGSTY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m701_MA1962_1 |
POU2F1::SOX2 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
13.85 |
20627 |
++++mATTWrCATmACAAWrg |
CYWTTGTKATGYWAATK++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m574_MA1646_1 |
OSR2 |
cluster_56 |
JASPAR_2022_vertebrates_CORE |
15 |
7.13 |
14808 |
+amaCAGAAGChr++ |
++YDGCTTCTGTKT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m785_MA0690_2 |
TBX21 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
9.40 |
7664 |
+++++++++AAGGTGTGAAA++ |
++tTTCACACCTt+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m76_MA0603_1 |
Arntl |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.64 |
1001 |
+++dgTCACGTGh+++++ |
+++++DCACGTGACH+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m400_MA1489_1 |
FOXN3 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
9.86 |
998 |
+++++++TTGTTTAC++++ |
++++GTAAACAA+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m753_MA0156_3 |
FEV |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.18 |
15106 |
++++YCACTTCCGGTY++++++ |
++++++rACCGGAAGTgr++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m156_MA0706_1 |
MEOX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.09 |
36834 |
+rsTaATtAwc++ |
++GWTAATTASY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m605_MA0640_2 |
ELF3 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.16 |
48502 |
++++AVCAGGAAGTGGRR++++ |
++++yyccACTTCCTgbt++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m496_MA1619_1 |
Ptf1A |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.90 |
7253 |
+rmaCAGCTGtky++ |
++RMACAGCTGTKY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m7_MA0040_1 |
Foxq1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
9.75 |
18 |
++++++WATAAACAWTA++ |
++tAwTGTTTATw++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m274_MA0846_1 |
FOXC2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.05 |
29281 |
++++wAwrTAAAYAww+++ |
+++WWTRTTTAYWTW++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m252_MA0816_1 |
Ascl2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.91 |
362 |
++arCAGCTGyy+++ |
+++RRCAGCTGYT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m638_MA0063_2 |
NKX2-5 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
7.15 |
5444 |
+++rmCACTCAAva++ |
++TBTTGAGTGKY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m473_MA1581_1 |
ZBTB6 |
cluster_83 |
JASPAR_2022_vertebrates_CORE |
13 |
8.37 |
2227 |
dysCTTGAGCCms |
SKGGCTCAAGSRH |
 |
 |
| JASPAR_2022_vertebrates_CORE_m385_MA1464_1 |
ARNT2 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
8.66 |
64620 |
++++rtCACGTGms++++ |
++++SKCACGTGAY++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m834_MA1567_2 |
Tbx6 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
8.05 |
9652 |
+++++++++rrGGTGTGAArw+ |
+WYTTCACACCYY+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m157_MA0709_1 |
Msx3 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.84 |
8871 |
++syAATTAr+++ |
+++YTAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m143_MA0083_3 |
SRF |
cluster_117 |
JASPAR_2022_vertebrates_CORE |
16 |
14.54 |
1913 |
TgmCCATATAWGGkcA |
TGMCCWTATATGGKCA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m735_MA1996_1 |
Nr1H2 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
7.34 |
8971 |
+++++++ywrAGGTCAyr++ |
++YRTGACCTYWR+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m106_MA0647_1 |
GRHL1 |
cluster_33 |
JASPAR_2022_vertebrates_CORE |
16 |
9.90 |
4033 |
aAAACCGGTTtd++++ |
++++HAAACCGGTTTT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m321_MA0907_1 |
HOXC13 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
9.84 |
1724 |
+DTTTTACGAGC+ |
+gCTCGTAAAAh+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m703_MA1964_1 |
SMAD2 |
cluster_64 |
JASPAR_2022_vertebrates_CORE |
12 |
7.48 |
3758 |
SYGTCTGGSR++ |
++ysCCAGACrs |
 |
 |
| JASPAR_2022_vertebrates_CORE_m758_MA0474_3 |
Erg |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.48 |
30061 |
+++rraCAGGAAGTgrs+++++ |
+++++SYCACTTCCTGTYY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m3_MA0019_1 |
Ddit3::Cebpa |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
8.08 |
39 |
+++GGKATTGCAYYY |
rrrTGCAATmcc+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m59_MA0519_1 |
Stat5a::Stat5b |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
9.24 |
16507 |
+++++TTCYKGGAAAY++ |
++rtTTCCmRGAA+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m236_MA0511_2 |
RUNX2 |
cluster_60 |
JASPAR_2022_vertebrates_CORE |
14 |
8.01 |
8974 |
+++TTGYGGTTW++ |
++wAACCRCAa+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m31_MA0139_1 |
CTCF |
cluster_76 |
JASPAR_2022_vertebrates_CORE |
35 |
11.93 |
913 |
++++++++++++++++ygrCCAsyAGrkGGCrsyr |
YRSYGCCMYCTRSTGGYCR++++++++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m1_MA0004_1 |
Arnt |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.62 |
20 |
++++++CACGTG++++++ |
++++++CACGTG++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m362_MA1135_1 |
FOSB::JUNB |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.81 |
1001 |
++++krTGAsTCAy++++++ |
++++++RTGASTCAYM++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m515_MA0893_2 |
GSX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.04 |
8512 |
++YTAATKRS+++ |
+++symATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m312_MA0894_1 |
HESX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.45 |
4293 |
+RYYAATTARC++ |
++gyTAATTrry+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m577_MA1650_1 |
ZBTB14 |
cluster_67 |
JASPAR_2022_vertebrates_CORE |
15 |
8.35 |
4547 |
+++ssCCGCGCACss |
SSGTGCGCGGSS+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m648_MA0743_2 |
SCRT1 |
cluster_37 |
JASPAR_2022_vertebrates_CORE |
16 |
9.42 |
60209 |
wawkCAACAGGTGgtw |
WACCACCTGTTGMWTW |
 |
 |
| JASPAR_2022_vertebrates_CORE_m740_MA2002_1 |
Zfp335 |
cluster_64 |
JASPAR_2022_vertebrates_CORE |
12 |
7.45 |
84 |
+RSTGCCTGAYY |
rrtCAGGCAsy+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m265_MA0838_1 |
CEBPG |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
10.16 |
22697 |
++rTTrCGCAAy+++ |
+++RTTGCGYAAY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m189_MA0737_1 |
GLIS3 |
cluster_46 |
JASPAR_2022_vertebrates_CORE |
16 |
14.58 |
981 |
+KACCCCCCACgAAG+ |
+CTTCGTGGGGGGTM+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m769_MA0526_4 |
USF2 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
11.27 |
19360 |
+++RRTCACGTGAYY+++ |
+++rrTCACGTGAyy+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m641_MA0712_2 |
OTX2 |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
7.40 |
120679 |
WTYTAATCCYHW |
wdrGGATTAraw |
 |
 |
| JASPAR_2022_vertebrates_CORE_m760_MA0489_2 |
Jun |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
7.94 |
94103 |
++++dvTGACTCAtyh++++ |
++++DRATGAGTCABH++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m4_MA0029_1 |
Mecom |
cluster_11 |
JASPAR_2022_vertebrates_CORE |
15 |
12.41 |
27 |
+TVTTATCTTRTCTT |
aAGAyAAGATAAba+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m372_MA1146_1 |
NR1H4::RXRA |
cluster_14 |
JASPAR_2022_vertebrates_CORE |
16 |
8.90 |
1001 |
grGkTCAtTGACCyy+ |
+RRGGTCAATGAMCYC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m651_MA0143_4 |
SOX2 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
6.94 |
85754 |
+++wwACAATGgww+++++ |
+++++WWCCATTGTWW+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m412_MA1505_1 |
HOXC8 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.51 |
13828 |
++gyAATTRc+++ |
+++GYAATTRC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m543_MA1708_1 |
ETV7 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
7.77 |
14072 |
++++YAYTTCCGSCY+++++++ |
+++++++rgscGGAARTr++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m771_MA0607_2 |
BHLHA15 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
9.52 |
13891 |
++aCCATATGGt+++ |
+++ACCATATGGT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m179_MA0727_1 |
NR3C2 |
cluster_19 |
JASPAR_2022_vertebrates_CORE |
17 |
13.16 |
58897 |
krGwACAywrTGTwCyc |
GRGWACAYWRTGTWCYM |
 |
 |
| JASPAR_2022_vertebrates_CORE_m578_MA1651_1 |
ZFP42 |
cluster_84 |
JASPAR_2022_vertebrates_CORE |
21 |
10.73 |
1752 |
swtccAArATGGCTGCcycsr |
YSGRGGCAGCCATYTTGGAWS |
 |
 |
| JASPAR_2022_vertebrates_CORE_m8_MA0051_1 |
IRF2 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
14.65 |
12 |
+++++++sGAAAGyGAAAsyrwwwm |
KWWWYRSTTTCRCTTTCS+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m395_MA1479_1 |
DMRTC2 |
cluster_26 |
JASPAR_2022_vertebrates_CORE |
12 |
8.50 |
1360 |
AATGTAKCAATT |
aaTTGmTACAtt |
 |
 |
| JASPAR_2022_vertebrates_CORE_m784_MA0685_2 |
SP4 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.95 |
999 |
+++CYCCKCCCC++++++ |
++++++gGGGMGGrG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m131_MA0672_1 |
NKX2-3 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
7.55 |
15942 |
+++asCACTTrav+++ |
+++BTYAAGTGST+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m61_MA0523_1 |
TCF7L2 |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
9.35 |
4188 |
++rrAswTCAAAGgva+++ |
+++TBCCTTTGAWSTYY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m840_MA0597_2 |
THAP1 |
cluster_49 |
JASPAR_2022_vertebrates_CORE |
12 |
7.73 |
5178 |
SSTGCCCTGCSS |
ssgcAGGGCAss |
 |
 |
| JASPAR_2022_vertebrates_CORE_m148_MA0692_1 |
TFEB |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.86 |
18927 |
++++RTCACGTGRY++++ |
++++ryCACGTGAy++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m839_MA1647_2 |
Prdm4 |
cluster_127 |
JASPAR_2022_vertebrates_CORE |
17 |
10.17 |
1181 |
rrgcCTtGAAACygyyr |
YRRCRGTTTCAAGGCYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m81_MA0614_1 |
Foxj2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.17 |
1000 |
+++++++rTAAACAa++++ |
++++TTGTTTAY+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m283_MA0857_1 |
Rarb |
cluster_47 |
JASPAR_2022_vertebrates_CORE |
19 |
18.29 |
2047 |
TGACCTTTTGACCTTT+++ |
+++aAAGGTCAAAAGGTCA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m354_MA1127_1 |
FOSB::JUN |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
8.90 |
1001 |
++++gaTGACGTCAy+++++ |
+++++RTGACGTCATC++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m444_MA1546_1 |
PAX3 |
cluster_91 |
JASPAR_2022_vertebrates_CORE |
16 |
10.24 |
1935 |
ysGTCACGsywATTAr |
YTAATWRSCGTGACSR |
 |
 |
| JASPAR_2022_vertebrates_CORE_m110_MA0046_2 |
HNF1A |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
11.01 |
19777 |
++++arTTAATsATTAACw |
WGTTAATSATTAAYT++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m636_MA0025_2 |
NFIL3 |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
8.01 |
22996 |
+wrTTATGyAAtaw+ |
+WTATTRCATAAYW+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m694_MA1955_1 |
FOXO1::ELK3 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.64 |
260 |
+rwmAACAGGAAGtr+++++++ |
+++++++YACTTCCTGTTKWY+ |
 |
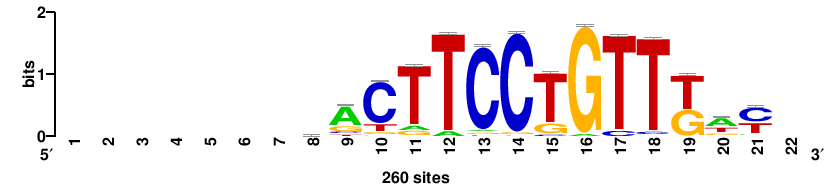 |
| JASPAR_2022_vertebrates_CORE_m294_MA0870_1 |
Sox1 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
14.51 |
2713 |
++++AACAATGTTATTGTT |
aACAATaaCATTGTt++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m505_MA1100_2 |
ASCL1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
6.51 |
6482 |
++RGCAGCTGYY+++ |
+++rrCAGCTGcy++ |
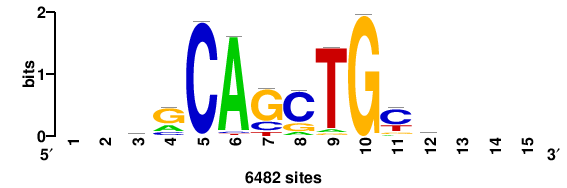 |
 |
| JASPAR_2022_vertebrates_CORE_m416_MA1509_1 |
IRF6 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
9.77 |
3553 |
+++++++++++ACCGAAACT+++++ |
+++++AGTTTCGGT+++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m520_MA0594_2 |
HOXA9 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
5.18 |
3608 |
++SRTTWAYGAY+ |
+rtCRTwAays++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m11_MA0069_1 |
PAX6 |
cluster_23 |
JASPAR_2022_vertebrates_CORE |
18 |
9.56 |
43 |
++MABTSAWGCGTRAA++ |
++TTyACGCwTsAvTk++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m242_MA0805_1 |
TBX1 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
8.89 |
14871 |
++++++++++AGGTGTGA++++ |
++++TCACACCT++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m607_MA0761_2 |
ETV1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.27 |
37339 |
+++rraCAGGAAGTgrr+++++ |
+++++YYCACTTCCTGTYY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m361_MA1134_1 |
FOS::JUNB |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
9.07 |
1001 |
++++krTGAsTCAymv++++ |
++++BKRTGASTCAYM++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m129_MA0671_1 |
NFIX |
cluster_49 |
JASPAR_2022_vertebrates_CORE |
12 |
5.41 |
59749 |
sgtGCCArv+++ |
+++BYTGGCACS |
 |
 |
| JASPAR_2022_vertebrates_CORE_m49_MA0483_1 |
Gfi1B |
cluster_55 |
JASPAR_2022_vertebrates_CORE |
12 |
9.37 |
1761 |
+aAATCwCwGCw |
WGCWGWGATTT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m693_MA1954_1 |
FOXO1::ELK1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.51 |
2398 |
+rwmAACAGGAAGtr+++++++ |
+++++++YACTTCCTGTTKWY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m459_MA1564_1 |
SP9 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
8.11 |
3844 |
++rcCmCGCCCmcy++++ |
++++RGKGGGCGKGGY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m338_MA1116_1 |
RBPJ |
cluster_44 |
JASPAR_2022_vertebrates_CORE |
17 |
7.79 |
2402 |
++++bsTGGGAAmg+++ |
+++CKTTCCCASV++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m644_MA0782_2 |
PKNOX1 |
cluster_34 |
JASPAR_2022_vertebrates_CORE |
17 |
8.55 |
39963 |
+twTGAgTGACAGctr+ |
+YAGCTGTCACTCAWA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m239_MA0802_1 |
TBR1 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
8.64 |
49124 |
++++++++++TTTCACACCT++ |
++AGGTGTGAAa++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m171_MA0722_1 |
VAX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.78 |
8033 |
++yTAATtAy+++ |
+++RTAATTAR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m737_MA1998_1 |
Prdm14 |
cluster_93 |
JASPAR_2022_vertebrates_CORE |
12 |
7.60 |
13841 |
hrGGTCTCTAac |
GTTAGAGACCYD |
 |
 |
| JASPAR_2022_vertebrates_CORE_m571_MA1643_1 |
NFIB |
cluster_28 |
JASPAR_2022_vertebrates_CORE |
21 |
13.04 |
4416 |
TDCTTGGCACMRTGCCAGGYA |
trcCTGGCAykgTGCCAagha |
 |
 |
| JASPAR_2022_vertebrates_CORE_m387_MA1468_1 |
ATOH7 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.51 |
1013 |
++RMCATATGKT+++ |
+++AmCATATGky++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m420_MA1515_1 |
KLF2 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
7.01 |
8161 |
+srCCmCrCCCw++++++ |
++++++WGGGYGKGGYS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m611_MA1103_2 |
FOXK2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
6.93 |
16679 |
+++++awGTAAACAar+++ |
+++YTTGTTTACWT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m687_MA1948_1 |
ETV5::HOXA2 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.06 |
486 |
+++++TAATKACWTCCGSY+++ |
+++rsCGGAwGTmATTA+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m398_MA1484_1 |
ETS2 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.24 |
21072 |
++++rACCGGAWGy++++++++ |
++++++++RCWTCCGGTY++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m311_MA0892_1 |
GSX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.40 |
1462 |
+mcymATTArh++ |
++DYTAATKRGK+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m528_MA0701_2 |
LHX9 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.61 |
8740 |
++CyAATTAr+++ |
+++YTAATTRG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m272_MA0845_1 |
FOXB1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.48 |
8178 |
++++wAwGTMAAyAt++++ |
++++ATRTTKACWTW++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m298_MA0874_1 |
Arx |
cluster_22 |
JASPAR_2022_vertebrates_CORE |
18 |
9.90 |
101 |
+WSMAYTAATTARYGSAC |
gtscryTAATTArtksw+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m273_MA0032_2 |
FOXC1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
8.23 |
23027 |
++++wAwRTMAAyAw++++ |
++++WTRTTKAYWTW++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m591_MA1730_1 |
ZNF708 |
cluster_57 |
JASPAR_2022_vertebrates_CORE |
15 |
10.49 |
849 |
RGGAGGYACAGCYGG |
ccrGCTGTrCCTccy |
 |
 |
| JASPAR_2022_vertebrates_CORE_m344_MA1121_1 |
TEAD2 |
cluster_2 |
JASPAR_2022_vertebrates_CORE |
19 |
7.71 |
542 |
+hcACATTCCwgsc+++++ |
+++++GSCWGGAATGTGD+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m153_MA0699_1 |
LBX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.10 |
3716 |
+rcyAATTArc++ |
++GYTAATTRGY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m536_MA0868_2 |
SOX8 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
7.05 |
4529 |
++agAACAATrg+++++++ |
+++++++CYATTGTTCT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m14_MA0072_1 |
RORA |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
12.08 |
36 |
+++wwwAwbTAGGTCAr+++ |
+++YTGACCTAVWTWWW+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m200_MA0758_1 |
E2F7 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
16.77 |
1053 |
+WTTTGGCGGGAAAA+++ |
+++tTTTCCCGCCAAAw+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m255_MA0817_1 |
BHLHE23 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
10.39 |
6808 |
+WAMCATATGKTT++ |
++aAmCATATGkTw+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m326_MA0036_3 |
GATA2 |
cluster_11 |
JASPAR_2022_vertebrates_CORE |
15 |
7.49 |
14213 |
RVAGATAAGRA++++ |
++++tyCTTATCTby |
 |
 |
| JASPAR_2022_vertebrates_CORE_m388_MA1101_2 |
BACH2 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
12.53 |
94294 |
AWABSATGASTCATGSTWT+ |
+awasCATGAsTCATsvtwt |
 |
 |
| JASPAR_2022_vertebrates_CORE_m95_MA0636_1 |
BHLHE41 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
10.73 |
9573 |
++++GTCACGTGAY++++ |
++++rTCACGTGAc++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m681_MA1942_1 |
ETV2::FOXI1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.22 |
3173 |
sgtaaACAGGAwRyr+++++++ |
+++++++YRYWTCCTGTTTACS |
 |
 |
| JASPAR_2022_vertebrates_CORE_m826_MA1511_2 |
KLF10 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.74 |
998 |
+++CCMCGCCCC++++++ |
++++++GGGGCGKGG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m47_MA0478_1 |
FOSL2 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
9.71 |
5318 |
+++++VTGASTCAYYM++++ |
++++krrTGAsTCAb+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m553_MA1718_1 |
ZNF8 |
cluster_70 |
JASPAR_2022_vertebrates_CORE |
20 |
17.44 |
2690 |
GTATrGATRTACCACAkTTT |
AAAMTGTGGTAYATCYATAC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m615_MA0482_2 |
GATA4 |
cluster_11 |
JASPAR_2022_vertebrates_CORE |
15 |
7.66 |
59014 |
WVAGATAAGGAA+++ |
+++ttcCTTATCTbw |
 |
 |
| JASPAR_2022_vertebrates_CORE_m21_MA0101_1 |
REL |
cluster_32 |
JASPAR_2022_vertebrates_CORE |
18 |
7.29 |
17 |
+++sGGrmwTTCC+++++ |
+++++GGAAWKYCCS+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m5_MA0030_1 |
FOXF2 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
10.28 |
28 |
++bmaasRTAAACAaw+++ |
+++WTTGTTTAYSTTKV++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m832_MA1563_2 |
SOX18 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
6.96 |
1357 |
++++aACAATdv+++++++ |
+++++++BHATTGTT++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m568_MA1640_1 |
MEIS2 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
8.31 |
20173 |
+RRYCATAAATCATTY+++ |
+++raaTGATTTATgryy+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m54_MA0501_1 |
MAF::NFE2 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
12.21 |
1090 |
+++++rTGACTCAGCArwwy |
RWWYTGCTGAGTCAY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m174_MA0726_1 |
VSX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.36 |
17977 |
++yTAATTAg+++ |
+++CTAATTAR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m285_MA0859_1 |
Rarg |
cluster_47 |
JASPAR_2022_vertebrates_CORE |
19 |
17.14 |
26801 |
+rRGGTCAAAAGGTCAa++ |
++TTGACCTTTTGACCYY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m428_MA1527_1 |
NFIC |
cluster_28 |
JASPAR_2022_vertebrates_CORE |
21 |
12.72 |
2966 |
++sTTGGCrysgTGCCArs++ |
++SYTGGCACSRYGCCAAS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m182_MA0730_1 |
RARA |
cluster_31 |
JASPAR_2022_vertebrates_CORE |
18 |
15.78 |
2020 |
+AGGTCAysyaRAGGTCA |
TGACCTYTRSRTGACCT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m768_MA0521_2 |
Tcf12 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.58 |
73995 |
+SVACAGCTGTBC++ |
++gvaCAGCTGtbs+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m794_MA0754_2 |
CUX1 |
cluster_35 |
JASPAR_2022_vertebrates_CORE |
10 |
8.66 |
12712 |
TAATCRATAm |
KTATYGATTA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m668_MA1929_1 |
CTCF |
cluster_76 |
JASPAR_2022_vertebrates_CORE |
35 |
12.70 |
6766 |
+cTGCagTkcchscwgyggCCasyAGrkGGCrsyv |
BRSYGCCMYCTRSTGGCCRCWGSDGGMACTGCAG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m707_MA1968_1 |
TFCP2 |
cluster_33 |
JASPAR_2022_vertebrates_CORE |
16 |
8.11 |
799 |
+RAACCGGTTT+++++ |
+++++aAACCGGTTy+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m763_MA0506_2 |
Nrf1 |
cluster_67 |
JASPAR_2022_vertebrates_CORE |
15 |
10.24 |
12729 |
YGCGCAKGCGCASRS |
sysTGCGCmTGCGcr |
 |
 |
| JASPAR_2022_vertebrates_CORE_m60_MA0520_1 |
Stat6 |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
9.75 |
1852 |
+RKTTCTSWRGAARWS++ |
++swyTTCywsaGAAmy+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m637_MA0502_2 |
NFYB |
cluster_17 |
JASPAR_2022_vertebrates_CORE |
13 |
8.58 |
7454 |
YTGGCCAATGRG+ |
+cyCATTGGCCar |
 |
 |
| JASPAR_2022_vertebrates_CORE_m138_MA0068_2 |
PAX4 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.56 |
617 |
++ctAATTAg+++ |
+++CTAATTAG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m204_MA0763_1 |
ETV3 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.06 |
304 |
+++++ACCGGAAGTg+++++++ |
+++++++CACTTCCGGT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m178_MA0113_3 |
NR3C1 |
cluster_19 |
JASPAR_2022_vertebrates_CORE |
17 |
13.30 |
27422 |
rrGwACAywrTGTwCym |
KRGWACAYWRTGTWCYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m72_MA0596_1 |
SREBF2 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
9.40 |
47 |
+++++++++rTGGgGTGAy+++ |
+++RTCACCCCAY+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m711_MA1972_1 |
ZFP14 |
cluster_80 |
JASPAR_2022_vertebrates_CORE |
15 |
10.53 |
999 |
GGAGGcmctGGAryG |
CRYTCCAGKGCCTCC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m41_MA0164_1 |
Nr2e3 |
cluster_110 |
JASPAR_2022_vertebrates_CORE |
8 |
8.34 |
23 |
+cAAGCTT |
AAGCTTG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m691_MA1952_1 |
FOXJ2::ELF1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.61 |
2625 |
+rtaaACmGGAAGwr+++++++ |
+++++++YWCTTCCKGTTTAY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m128_MA0670_1 |
NFIA |
cluster_49 |
JASPAR_2022_vertebrates_CORE |
12 |
8.44 |
98253 |
ggTGCCAArw++ |
++WYTTGGCACC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m535_MA0867_2 |
SOX4 |
cluster_8 |
JASPAR_2022_vertebrates_CORE |
19 |
7.76 |
12829 |
+++gaACAAwGrv++++++ |
++++++BYCWTTGTTC+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m666_MA1684_1 |
Foxn1 |
cluster_110 |
JASPAR_2022_vertebrates_CORE |
8 |
6.59 |
5235 |
rGAmGC++ |
++GCKTCY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m79_MA0610_1 |
DMRT3 |
cluster_26 |
JASPAR_2022_vertebrates_CORE |
12 |
7.79 |
1001 |
ATTGATACATT+ |
+aaTGTATCaat |
 |
 |
| JASPAR_2022_vertebrates_CORE_m62_MA0527_1 |
ZBTB33 |
cluster_69 |
JASPAR_2022_vertebrates_CORE |
15 |
9.94 |
705 |
yTCTCGCGakayyts |
SARRTMTCGCGAGAR |
 |
 |
| JASPAR_2022_vertebrates_CORE_m201_MA0760_1 |
ERF |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.94 |
304 |
+++++ACCGGAArTr+++++++ |
+++++++YAYTTCCGGT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m351_MA1155_1 |
ZSCAN4 |
cluster_132 |
JASPAR_2022_vertebrates_CORE |
15 |
14.65 |
2144 |
tGCACACmCTGaAAa |
TTTTCAGKGTGTGCA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m689_MA1950_1 |
FLI1::FOXI1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.37 |
3032 |
+rtaAACAGGAwryd+++++++ |
+++++++HRYWTCCTGTTTAY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m222_MA0786_1 |
POU3F1 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
8.29 |
4173 |
+++WMATTWRCATAW++++++ |
++++++wTATGywAATkw+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m557_MA1722_1 |
ZSCAN31 |
cluster_133 |
JASPAR_2022_vertebrates_CORE |
19 |
17.20 |
9868 |
GCATAAyTGCCCTGCkkCc |
GGMMGCAGGGCARTTATGC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m573_MA1645_1 |
NKX2-2 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
7.45 |
26023 |
+hawCCACTCAAvwa+ |
+TWBTTGAGTGGWTD+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m748_MA0080_6 |
Spi1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.29 |
105198 |
+rraAAGAGGAAGTGgra++++ |
++++TYCCACTTCCTCTTTYY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m194_MA0749_1 |
ZBED1 |
cluster_69 |
JASPAR_2022_vertebrates_CORE |
15 |
13.21 |
6111 |
cTrTCGCGACATr++ |
++YATGTCGCGAYAG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m484_MA1597_1 |
ZNF528 |
cluster_103 |
JASPAR_2022_vertebrates_CORE |
17 |
15.15 |
616 |
cccAGGGAAGcCATTtC |
GAAATGGCTTCCCTGGG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m392_MA1474_1 |
CREB3L4 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
7.56 |
6369 |
+++GRTGACGTGGCR+++ |
+++ygCCACGTCAyc+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m619_MA0114_4 |
HNF4A |
cluster_2 |
JASPAR_2022_vertebrates_CORE |
19 |
8.45 |
46504 |
kyCAAAGTCCArr++++++ |
++++++YYTGGACTTTGRM |
 |
 |
| JASPAR_2022_vertebrates_CORE_m465_MA1572_1 |
TGIF2LY |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.73 |
7196 |
+TGACAGCTGTCA++ |
++TGACAgcTGTCA+ |
 |
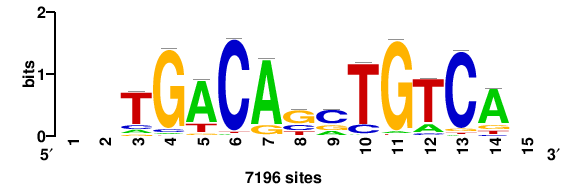 |
| JASPAR_2022_vertebrates_CORE_m656_MA0769_2 |
TCF7 |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
7.59 |
14994 |
++++ASWTCAAAGRM++++ |
++++kyCTTTGAwst++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m154_MA0704_1 |
Lhx4 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.77 |
4025 |
++YTAATTAR+++ |
+++yTAATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m744_MA0050_3 |
Irf1 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
9.49 |
15640 |
+++++amtGAAAcTGAAAsy+++++ |
+++++RSTTTCAGTTTCAKT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m240_MA0803_1 |
TBX15 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
9.22 |
11937 |
++++++++++AGGTGTGA++++ |
++++TCACACCT++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m195_MA0751_1 |
ZIC4 |
cluster_46 |
JASPAR_2022_vertebrates_CORE |
16 |
9.43 |
7525 |
+krCCCCCyGykGyGh |
DCRCMRCRGGGGGYM+ |
 |
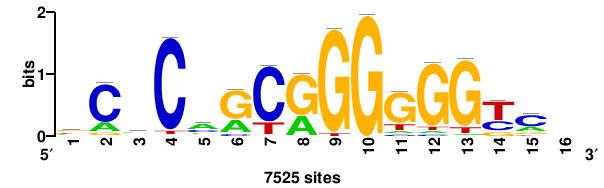 |
| JASPAR_2022_vertebrates_CORE_m602_MA0162_4 |
EGR1 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
8.32 |
30598 |
+SKGCGTGGGCGKGG+++ |
+++ccmCGCCCACgCms+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m333_MA0060_3 |
NFYA |
cluster_17 |
JASPAR_2022_vertebrates_CORE |
13 |
8.42 |
4825 |
++rrCCAATCAgm |
KCTGATTGGYY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m121_MA0661_1 |
MEOX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.67 |
4038 |
+rsTAATTAmc++ |
++GKTAATTASY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m217_MA0779_1 |
PAX1 |
cluster_23 |
JASPAR_2022_vertebrates_CORE |
18 |
13.62 |
6849 |
+sGTCACGCwTSAsTGmm |
KKCASTSAWGCGTGACS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m477_MA1587_1 |
ZNF135 |
cluster_65 |
JASPAR_2022_vertebrates_CORE |
16 |
12.43 |
948 |
+cCTCGACCTCCyrr+ |
+YYRGGAGGTCGAGG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m254_MA0464_2 |
BHLHE40 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
8.80 |
13094 |
++++GKCACGTGMB++++ |
++++vkCACGTGmc++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m658_MA0809_2 |
TEAD4 |
cluster_2 |
JASPAR_2022_vertebrates_CORE |
19 |
7.95 |
78086 |
+SCTGGAATGTGR++++++ |
++++++ycACATTCCAgs+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m36_MA0065_2 |
Pparg::Rxra |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
8.08 |
864 |
++strGGgcArAGGkcA++ |
++TGMCCTYTGCCCYAS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m563_MA1635_1 |
BHLHE22 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.42 |
18356 |
++CSCAGCTGSG+++ |
+++csCAGCTGsg++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m101_MA0645_1 |
ETV6 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.56 |
22662 |
+++++YACTTCCGSK+++++++ |
+++++++msCGGAAGTr+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m522_MA0904_2 |
HOXB5 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.62 |
4160 |
++sTMATKAs+++ |
+++STMATKAS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m796_MA0759_2 |
ELK3 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.83 |
21909 |
+++++aCCGGAAGTrs++++++ |
++++++SYACTTCCGGT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m808_MA0837_2 |
CEBPE |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
11.31 |
15314 |
+KATTGCGCAATM++ |
++kATTGCGCAATm+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m624_MA0039_4 |
KLF4 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
7.61 |
55817 |
+SDGGGTGGGGSS+++++ |
+++++ssCCCCACCChs+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m775_MA0633_2 |
Twist2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.23 |
49629 |
++vvCAGCTGbb+++ |
+++VVCAGCTGBB++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m526_MA0913_2 |
HOXD9 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
8.68 |
26608 |
+++DTTTTATKRC++ |
++GymATAAAAh+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m290_MA0862_1 |
GMEB2 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
6.85 |
457 |
++++++ttACGTAa++++++ |
++++++TTACGTAA++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m369_MA1143_1 |
FOSL1::JUND |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
5.13 |
1000 |
+++++rTGACgymay+++++ |
+++++RTKRCGTCAY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m533_MA0682_2 |
PITX1 |
cluster_29 |
JASPAR_2022_vertebrates_CORE |
12 |
7.70 |
15811 |
++yTAATCCc++ |
++GGGATTAR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m233_MA0796_1 |
TGIF1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
15.44 |
13670 |
+TGACAGCTGTCA++ |
++TGACAGCTGTCA+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m442_MA1544_1 |
OVOL1 |
cluster_48 |
JASPAR_2022_vertebrates_CORE |
17 |
10.51 |
24189 |
arwACCGTTAttys+++ |
+++SRAATAACGGTWYT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m667_MA1928_1 |
BNC2 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
7.59 |
5672 |
++++WRTGACTCAYWW++++ |
++++wwrTGAGTCAyw++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m159_MA0124_2 |
Nkx3-1 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
7.81 |
10383 |
+++rcCACTTAa++++ |
++++TTAAGTGGY+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m402_MA1494_1 |
HNF4A |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
14.52 |
896 |
+++KYGACCTTTGGACYY+ |
+rrGTCCAAAGGTCRm+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m628_MA0496_3 |
MAFK |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.81 |
51734 |
+++tgmTGACTCAGCmww++ |
++WWKGCTGAGTCAKCA+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m383_MA1421_1 |
TCF7L1 |
cluster_4 |
JASPAR_2022_vertebrates_CORE |
19 |
11.63 |
203 |
++CCTTTGATCTTT+++++ |
+++++AAAGATCAAAGG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m331_MA1109_1 |
NEUROD1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.38 |
2282 |
SYRCCATCTGTYC++ |
++graCAGATGGyrs |
 |
 |
| JASPAR_2022_vertebrates_CORE_m542_MA1707_1 |
DMRTA1 |
cluster_26 |
JASPAR_2022_vertebrates_CORE |
12 |
7.61 |
4949 |
AWTGTAWCAATT |
aattGwTACAwt |
 |
 |
| JASPAR_2022_vertebrates_CORE_m589_MA1728_1 |
ZNF549 |
cluster_86 |
JASPAR_2022_vertebrates_CORE |
12 |
7.80 |
4827 |
rsTGCTGCCCwr |
YWGGGCAGCASY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m401_MA1493_1 |
HES6 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
9.81 |
533 |
++++GGCACGTGTy++++ |
++++RACACGTGCC++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m679_MA1940_1 |
ETV2::DRGX |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.02 |
2416 |
+++++rsCGGAWryaATTA+++ |
+++TAATTRYWTCCGSY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m663_MA0748_2 |
YY2 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
7.54 |
5704 |
+SGCCGCCATSK++++++ |
++++++msATGGCGGcs+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m12_MA0070_1 |
PBX1 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
10.15 |
18 |
+++WWTGATTGATGD++++ |
++++hcATCAATCAww+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m203_MA0762_1 |
ETV2 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
10.16 |
2202 |
++++aACCGGAARYr+++++++ |
+++++++YRYTTCCGGTT++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m352_MA0099_3 |
FOS::JUN |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.14 |
1000 |
++++KRTGACTCAY++++++ |
++++++rTGAGTCAym++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m132_MA0673_1 |
NKX2-8 |
cluster_54 |
JASPAR_2022_vertebrates_CORE |
16 |
6.26 |
12324 |
++++sCACTYsAr+++ |
+++YTSRAGTGS++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m599_MA1102_2 |
CTCFL |
cluster_62 |
JASPAR_2022_vertebrates_CORE |
14 |
8.67 |
18037 |
+rsCAGGGGGCgs+ |
+SCGCCCCCTGSY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m518_MA0900_2 |
HOXA2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.50 |
60585 |
++STAATKAS+++ |
+++sTMATTAs++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m463_MA1570_1 |
TFAP4 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.27 |
915 |
++AMCAYRTGDT+++ |
+++ahCAyrTGkt++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m498_MA1621_1 |
Rbpjl |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.55 |
9652 |
svmaCACCTGtsyc+ |
+GRSACAGGTGTKBS |
 |
 |
| JASPAR_2022_vertebrates_CORE_m476_MA1585_1 |
ZKSCAN1 |
cluster_66 |
JASPAR_2022_vertebrates_CORE |
11 |
9.35 |
6127 |
AyAGTAGGtg+ |
+CACCTACTRT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m318_MA0903_1 |
HOXB3 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
5.33 |
15247 |
+asymATTArc++ |
++GYTAATKRST+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m489_MA1604_1 |
Ebf2 |
cluster_7 |
JASPAR_2022_vertebrates_CORE |
15 |
8.71 |
26261 |
++kyCCCAAGGGAvh |
DBTCCCTTGGGRM++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m487_MA1602_1 |
ZSCAN29 |
cluster_15 |
JASPAR_2022_vertebrates_CORE |
12 |
10.29 |
999 |
SCYGYGTAGACR |
yGTCTACrCrGs |
 |
 |
| JASPAR_2022_vertebrates_CORE_m835_MA1573_2 |
Thap11 |
cluster_121 |
JASPAR_2022_vertebrates_CORE |
24 |
12.28 |
660 |
++++++++++ACTACAabTCCCAg |
CTGGGAVTTGTAGT++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m314_MA0896_1 |
Hmx1 |
cluster_22 |
JASPAR_2022_vertebrates_CORE |
18 |
9.43 |
101 |
+AWBCWTTAATTGCTBSK |
msvAGCAATTAAwgvwt+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m176_MA0141_3 |
ESRRB |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
11.32 |
2302 |
+++++++TCAAGGTCAww++ |
++WWTGACCTTGA+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m149_MA0694_1 |
ZBTB7B |
cluster_36 |
JASPAR_2022_vertebrates_CORE |
14 |
9.68 |
3434 |
+rCGACCMCCgaa+ |
+TTCGGKGGTCGY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m503_MA1628_1 |
Zic1::Zic2 |
cluster_56 |
JASPAR_2022_vertebrates_CORE |
15 |
8.20 |
9892 |
++cvCAGCAGGsr++ |
++YSCCTGCTGBG++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m722_MA1983_1 |
ZNF582 |
cluster_130 |
JASPAR_2022_vertebrates_CORE |
21 |
16.15 |
293 |
tcTgTTACTTGCAGCCaAAwg |
CWTTTGGCTGCAAGTAACAGA |
 |
 |
| JASPAR_2022_vertebrates_CORE_m704_MA1965_1 |
SP5 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
8.39 |
20696 |
+++cyCCTCCCcs+++++ |
+++++SGGGGAGGRG+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m126_MA0667_1 |
MYF6 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
10.11 |
610 |
++AACArCTGTY+++ |
+++RACAGYTGTT++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m85_MA0621_1 |
mix-a |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
8.67 |
1001 |
+MCTAATTAAYT+ |
+artTAATTAgk+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m509_MA0864_2 |
E2F2 |
cluster_53 |
JASPAR_2022_vertebrates_CORE |
18 |
9.09 |
6905 |
rwtttGGCGCCawwwy++ |
++RWWWTGGCGCCAAAWY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m621_MA0901_2 |
HOXB13 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
9.09 |
78714 |
+WWWTTTTATTGGWW |
wwcCAATAAAAwww+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m702_MA1963_1 |
SATB1 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
8.84 |
34224 |
WWDTTATTAGWWW++ |
++wwwCTAATAAhww |
 |
 |
| JASPAR_2022_vertebrates_CORE_m186_MA0733_1 |
EGR4 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.63 |
9141 |
+yymCGCCCACGCAhwt+ |
+AWDTGCGTGGGCGKRR+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m367_MA1141_1 |
FOS::JUND |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
9.22 |
1001 |
+++BKRTGACTCAYMM++++ |
++++kkrTGAGTCAymv+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m370_MA1144_1 |
FOSL2::JUND |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.98 |
1001 |
++++krTGAsTCAy++++++ |
++++++RTGASTCAYM++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m774_MA0625_2 |
NFATC3 |
cluster_44 |
JASPAR_2022_vertebrates_CORE |
17 |
5.82 |
9016 |
+++++ryGGAAamw+++ |
+++WKTTTCCRY+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m122_MA0662_1 |
MIXL1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.98 |
34728 |
+wyyAATTArm++ |
++KYTAATTRRW+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m595_MA0465_2 |
CDX2 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
7.94 |
32026 |
+adGCAATAAAaw++ |
++WTTTTATTGCHT+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m709_MA1970_1 |
TRPS1 |
cluster_11 |
JASPAR_2022_vertebrates_CORE |
15 |
7.21 |
18283 |
WMAGATAAGAWW+++ |
+++wwTCTTATCTkw |
 |
 |
| JASPAR_2022_vertebrates_CORE_m836_MA1601_2 |
ZNF75D |
cluster_44 |
JASPAR_2022_vertebrates_CORE |
17 |
11.21 |
999 |
+++++GTGGGAAAgCCT |
AGGCTTTCCCAC+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m223_MA0787_1 |
POU3F2 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
9.55 |
2377 |
+++TMATTWGCATAW++++++ |
++++++wTATGCwAATka+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m767_MA0516_3 |
SP2 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.51 |
998 |
+++CCCCGCCCC++++++ |
++++++GGGGCGGGg+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m243_MA0806_1 |
TBX4 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
6.86 |
33781 |
++++++++++AGGTGTgA++++ |
++++TCACACCT++++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m399_MA1485_1 |
FERD3L |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
12.52 |
41187 |
GTRACAGCTGKYRC+ |
+GyrmCAGCTGTyAC |
 |
 |
| JASPAR_2022_vertebrates_CORE_m376_MA1150_1 |
RORB |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
6.51 |
1001 |
++++++RTGACCYAAWT+++ |
+++AwttrGGTCAy++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m551_MA1716_1 |
ZNF76 |
cluster_63 |
JASPAR_2022_vertebrates_CORE |
21 |
13.32 |
2947 |
mytwCCCAyAATGCAyygCgc |
GCGCRRTGCATTRTGGGWARK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m642_MA1113_2 |
PBX2 |
cluster_41 |
JASPAR_2022_vertebrates_CORE |
19 |
8.29 |
29596 |
++rhcATAAATCAtw++++ |
++++WATGATTTATGDY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m256_MA0820_1 |
FIGLA |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.20 |
2307 |
++WMCAGGTGKW+++ |
+++wmCACCTGkw++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m558_MA1723_1 |
PRDM9 |
cluster_78 |
JASPAR_2022_vertebrates_CORE |
27 |
11.56 |
999 |
+rGkGGgmAgGGrGGmrrmArvarr++ |
++YYTBYTKYYKCCYCCCTKCCCMCY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m80_MA0613_1 |
FOXG1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
9.25 |
1000 |
+++++++rTAAACAw++++ |
++++WTGTTTAY+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m221_MA0785_1 |
POU2F1 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
8.35 |
15608 |
++++AATTWGCATAWT+++++ |
+++++awTATGcwAATt++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m564_MA1636_1 |
CEBPG |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
9.52 |
37316 |
adATGATGCAATmwt |
AWKATTGCATCATHT |
 |
 |
| JASPAR_2022_vertebrates_CORE_m545_MA1710_1 |
ZNF257 |
cluster_74 |
JASPAR_2022_vertebrates_CORE |
14 |
9.53 |
998 |
++swGAGGcrAGrg |
CYCTYGCCTCWS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m790_MA0723_2 |
VAX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
4.86 |
22739 |
++YTAATTRS+++ |
+++syaATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m20_MA0092_1 |
Hand1::Tcf3 |
cluster_64 |
JASPAR_2022_vertebrates_CORE |
12 |
7.03 |
29 |
+srTCTGGmwt+ |
+AWKCCAGAYS+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m632_MA0499_2 |
MYOD1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
8.14 |
34300 |
+cwgCACCTGTymy+ |
+RKRACAGGTGCWG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m612_MA0481_3 |
FOXP1 |
cluster_10 |
JASPAR_2022_vertebrates_CORE |
19 |
6.95 |
74718 |
+++++awGTAAACAww+++ |
+++WWTGTTTACWT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m23_MA0108_2 |
TBP |
cluster_95 |
JASPAR_2022_vertebrates_CORE |
21 |
6.99 |
389 |
++++sTATAWAwrssssss++ |
++SSSSSSYWTWTATAS++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m592_MA1731_1 |
ZNF768 |
cluster_104 |
JASPAR_2022_vertebrates_CORE |
18 |
10.05 |
999 |
yygcyyrsCcTCTCTGdk |
MHCAGAGAGGSYRRGCRR |
 |
 |
| JASPAR_2022_vertebrates_CORE_m257_MA0822_1 |
HES7 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
13.71 |
2055 |
+++YGGCACGTGCCR+++ |
+++yGGCACGTGCCr+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m756_MA0466_3 |
CEBPB |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
11.41 |
16055 |
+kATTGCGCAATm++ |
++KATTGCGCAATM+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m613_MA0062_3 |
GABPA |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
8.16 |
65479 |
+++RRACAGGAAGTGRR+++++ |
+++++yycACTTCCTGtyy+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m770_MA0606_2 |
Nfat5 |
cluster_44 |
JASPAR_2022_vertebrates_CORE |
17 |
7.94 |
356 |
+++amATGGAAAAtk++ |
++MATTTTCCATKT+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m540_MA0753_2 |
ZNF740 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
10.53 |
2437 |
GTGGGGGGGGYRG+++++ |
+++++cyrCCCCCCCCAc |
 |
 |
| JASPAR_2022_vertebrates_CORE_m75_MA0602_1 |
Arid5a |
cluster_101 |
JASPAR_2022_vertebrates_CORE |
14 |
7.19 |
101 |
cyAATATTgmdadw |
WHTHKCAATATTRG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m237_MA0800_1 |
EOMES |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
8.60 |
20891 |
+++++++++aAGGTGTGAaawh |
DWTTTCACACCTT+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m661_MA1123_2 |
TWIST1 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
7.79 |
27333 |
+AVACATCTGGWWT+ |
+awwCCAGATGtbt+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m752_MA0152_2 |
Nfatc2 |
cluster_44 |
JASPAR_2022_vertebrates_CORE |
17 |
8.07 |
3239 |
AWWKTTTCCATTKCHW+ |
+wdgmaATGGAAAmwwt |
 |
 |
| JASPAR_2022_vertebrates_CORE_m822_MA1483_2 |
ELF2 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
9.26 |
146401 |
+++awmCCGGAAGTr+++++++ |
+++++++YACTTCCGGKWT+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m777_MA0644_2 |
ESX1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.94 |
22850 |
++yyAATTAv+++ |
+++BTAATTRR++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m548_MA1713_1 |
ZNF610 |
cluster_77 |
JASPAR_2022_vertebrates_CORE |
16 |
9.93 |
2386 |
sscGCCGCTCCsss++ |
++SSSGGAGCGGCGSS |
 |
 |
| JASPAR_2022_vertebrates_CORE_m151_MA0696_1 |
ZIC1 |
cluster_46 |
JASPAR_2022_vertebrates_CORE |
16 |
13.01 |
39709 |
+CRCAGCRGGGGGTC+ |
+gACCCCCyGCTGyG+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m423_MA1519_1 |
LHX5 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.65 |
8750 |
++sYAATTAa+++ |
+++TTAATTRS++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m133_MA0674_1 |
NKX6-1 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.37 |
2464 |
++TTAATKRS+++ |
+++sYmATTAa++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m16_MA0074_1 |
RXRA::VDR |
cluster_61 |
JASPAR_2022_vertebrates_CORE |
17 |
14.18 |
10 |
+RGGTCAwcGrGTTCA+ |
+TGAACYCGWTGACCY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m373_MA1147_1 |
NR4A2::RXRA |
cluster_14 |
JASPAR_2022_vertebrates_CORE |
16 |
9.93 |
1001 |
rrGGTCrtTGACCyc+ |
+GRGGTCAAYGACCYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m391_MA1473_1 |
CDX4 |
cluster_45 |
JASPAR_2022_vertebrates_CORE |
15 |
9.92 |
36855 |
++gGymATAAAac++ |
++GTTTTATKRCC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m429_MA1528_1 |
NFIX |
cluster_28 |
JASPAR_2022_vertebrates_CORE |
21 |
11.08 |
29185 |
++gtTGGCrysgTGCCArs++ |
++SYTGGCACSRYGCCAAC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m838_MA1633_2 |
BACH1 |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
9.78 |
48991 |
+++++ATGACTCAt++++++ |
++++++ATGAGTCAT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m319_MA0905_1 |
HOXC10 |
cluster_12 |
JASPAR_2022_vertebrates_CORE |
13 |
7.66 |
5879 |
++WTTTWAYGAY+ |
+rTCrTwAAaw++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m304_MA0883_1 |
Dmbx1 |
cluster_108 |
JASPAR_2022_vertebrates_CORE |
17 |
9.00 |
101 |
wdrwmmGGATTArkbww |
WWVMYTAATCCKKWYHW |
 |
 |
| JASPAR_2022_vertebrates_CORE_m393_MA1475_1 |
CREB3L4 |
cluster_18 |
JASPAR_2022_vertebrates_CORE |
20 |
7.72 |
1851 |
++++grTGACGTCAyc++++ |
++++GRTGACGTCAYC++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m225_MA0789_1 |
POU3F4 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
9.27 |
14420 |
+++++ATTWGCATA+++++++ |
+++++++TATGCwAAT+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m645_MA0627_2 |
POU2F3 |
cluster_24 |
JASPAR_2022_vertebrates_CORE |
21 |
9.29 |
56029 |
+++WWATTTGCATATW+++++ |
+++++watATGCAAATww+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m164_MA0715_1 |
PROP1 |
cluster_38 |
JASPAR_2022_vertebrates_CORE |
16 |
10.48 |
673 |
++TAATywAATTA+++ |
+++TAATTWRATTA++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m114_MA0653_1 |
IRF9 |
cluster_42 |
JASPAR_2022_vertebrates_CORE |
25 |
17.21 |
4314 |
+++++RGTTTCGGTTTCGWT+++++ |
+++++AwCGAAACCGAAACy+++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m406_MA1499_1 |
HOXB4 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
7.20 |
11877 |
++YTAATKAY+++ |
+++rTmATTAr++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m268_MA0117_2 |
Mafb |
cluster_51 |
JASPAR_2022_vertebrates_CORE |
16 |
7.86 |
3127 |
+++awwwTGCTGACd+ |
+HGTCAGCAWWWT+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m28_MA0125_1 |
Nobox |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.64 |
38 |
+++TAATTrsy++ |
++RSYAATTA+++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m421_MA1516_1 |
KLF3 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
9.96 |
49050 |
+WGGGCGYGGYC++++++ |
++++++grCCrCGCCCw+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m270_MA0843_1 |
TEF |
cluster_5 |
JASPAR_2022_vertebrates_CORE |
15 |
10.11 |
4064 |
+KRTTAYGTAAYR++ |
++yrTTACrTAAym+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m357_MA1130_1 |
FOSL2::JUN |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
8.41 |
1001 |
++++KRTGACTCAYMM++++ |
++++kkrTGAGTCAym++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m207_MA0771_1 |
HSF4 |
cluster_16 |
JASPAR_2022_vertebrates_CORE |
15 |
9.45 |
18902 |
++GAAYRTTCYAGAA |
TTCtrGAAyrTTC++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m66_MA0137_3 |
STAT1 |
cluster_43 |
JASPAR_2022_vertebrates_CORE |
18 |
10.11 |
3629 |
++++tTTCyrGGAAa+++ |
+++TTTCCYRGAAA++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m309_MA0890_1 |
GBX2 |
cluster_3 |
JASPAR_2022_vertebrates_CORE |
13 |
6.95 |
29276 |
+rsyAATTArs++ |
++SYTAATTRSY+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m177_MA0017_2 |
NR2F1 |
cluster_39 |
JASPAR_2022_vertebrates_CORE |
20 |
9.25 |
48922 |
+++++++CYYKTGACCYYTG |
carRGGTCAmrrG+++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m494_MA1616_1 |
Prdm15 |
cluster_109 |
JASPAR_2022_vertebrates_CORE |
15 |
9.17 |
2237 |
rrgAAAACCTGGAry |
RYTCCAGGTTTTCYY |
 |
 |
| JASPAR_2022_vertebrates_CORE_m187_MA0735_1 |
GLIS1 |
cluster_46 |
JASPAR_2022_vertebrates_CORE |
16 |
17.58 |
10139 |
mGACCCCCCACGWwGc |
GCWWCGTGGGGGGTCK |
 |
 |
| JASPAR_2022_vertebrates_CORE_m146_MA0689_1 |
TBX20 |
cluster_13 |
JASPAR_2022_vertebrates_CORE |
22 |
9.60 |
811 |
+++++++++YTTCACACCTW++ |
++wAGGTGTGAAr+++++++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m583_MA1656_1 |
ZNF449 |
cluster_92 |
JASPAR_2022_vertebrates_CORE |
14 |
9.18 |
7660 |
cvaAGCCCAACcas |
STGGTTGGGCTTBG |
 |
 |
| JASPAR_2022_vertebrates_CORE_m368_MA1142_1 |
FOSL1::JUND |
cluster_1 |
JASPAR_2022_vertebrates_CORE |
20 |
5.17 |
1001 |
++++rrTGActmAt++++++ |
++++++ATKAGTCAYY++++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m837_MA1630_2 |
ZNF281 |
cluster_27 |
JASPAR_2022_vertebrates_CORE |
18 |
11.79 |
999 |
+YBCCCCTCCCCC+++++ |
+++++GGGGGAGGGGvr+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m127_MA0669_1 |
NEUROG2 |
cluster_6 |
JASPAR_2022_vertebrates_CORE |
15 |
10.05 |
746 |
++RACATATGTY+++ |
+++raCATATGtY++ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m695_MA1956_1 |
FOXO1::FLI1 |
cluster_9 |
JASPAR_2022_vertebrates_CORE |
22 |
12.78 |
380 |
+GwmAACAGGAAGTk+++++++ |
+++++++MACTTCCTGTTKWC+ |
 |
 |
| JASPAR_2022_vertebrates_CORE_m807_MA0831_3 |
TFE3 |
cluster_21 |
JASPAR_2022_vertebrates_CORE |
18 |
10.67 |
23570 |
++++rTCACGTGAy++++ |
++++RTCACGTGAY++++ |
 |
 |